M8 · Migration from siteBayes2
JoonHo Lee
2026-05-10
Source:vignettes/m8-migration-from-siteBayes2.Rmd
m8-migration-from-siteBayes2.RmdAbstract
For users of the internal siteBayes2 package who
need to migrate scripts to multisiteDGP. The new package keeps the
data-generating process and drops the Stan-based fitting, so the
migration is mostly a rename map plus a removal list. You leave
with eight worked before/after patterns, a function rename table,
an explicit deprecated-functions table that points to downstream
Bayesian packages, and the boundary check confirming
siteBayes2 exports are absent from the multisiteDGP
namespace.
1. Scope of the migration
multisiteDGP carries forward the data-generating process from
siteBayes2 and leaves Bayesian fitting to downstream
packages. Two practical consequences for an existing script:
-
Renamed surface.
sim_multisite_data(),gen_priorG(),sim_sitesize_withinvar(),sim_observed_effects(),get_shrinkage_factor()and friends becomesim_multisite(),gen_effects(),gen_site_sizes(),align_rank_corr()plusgen_observations(), andmean_shrinkage()— same idea, cleaner separation across the four generative layers (Lee et al., 2025). -
Removed surface.
fit_rubin_normal(),lm_stan(),data_prep_rubin_normal()and theinst/stan/model objects do not move. multisiteDGP installs without a C++ compiler. Fit withmetafor,baggr, orrstandirectly via the adapters in M6: Adapters and downstream packages.
Sections 2 and 3 give the rename and removal tables. Sections 4 through 11 walk through eight before/after patterns (MV1–MV8) — each runs as printed. Sections 12 and 13 cover the boundary check and where to read next.
2. Function rename map
The canonical translation from siteBayes2 to
multisiteDGP. Every target name in the right column appears in
NAMESPACE; verify with
grep '^export' multisiteDGP-R-package/NAMESPACE.
| siteBayes2 | multisiteDGP | Notes |
|---|---|---|
sim_multisite_data() |
sim_multisite() |
Front door; refactored into the four-layer pipeline |
gen_priorG() |
gen_effects() |
Layer 1; selects from eight built-in shapes |
gen_priorG2() |
gen_effects() with
formula, beta, data
|
Same function; covariate-aware path is an argument set, not a separate function |
sim_sitesize_withinvar() |
gen_site_sizes() |
Layer 2; engine choice replaces the implicit JEBS algorithm |
sim_observed_effects() |
align_rank_corr()
+ gen_observations()
|
Layer 3 (dependence) and Layer 4 (observation) are now explicit |
get_shrinkage_factor() |
mean_shrinkage() |
Vectorized over nj_mean; returns the mean-shrinkage
scalar diagnostic |
precision_dependence = TRUE |
sim_multisite()
with dependence = "rank" and rank_corr
|
Three named injection methods replace the boolean flag |
precision_dependence = FALSE |
sim_multisite()
with dependence = "none" (default) |
Same intent: precision and effect are sampled independently |
3. Deprecated functions — no equivalent in multisiteDGP
The Stan-based fitting surface is intentionally absent. There is no shim and no compatibility layer in multisiteDGP itself; downstream packages are the destination.
| siteBayes2 | Where to go now | Why removed |
|---|---|---|
fit_rubin_normal() |
metafor::rma() after as_metafor();
baggr::baggr() after as_baggr()
|
Frequentist and Bayesian random-effects fitting are first-class in
metafor/baggr
|
lm_stan() |
rstanarm::stan_lm(), brms::brm()
|
General-purpose Bayesian linear regression belongs in a general-purpose package |
data_prep_rubin_normal() |
as_metafor(), as_baggr()
|
The adapter renames effect, SE, and site columns to the target convention |
prior_rubin_normal() |
baggr::baggr() argument prior_hypermean,
prior_hypersd
|
Prior elicitation is the fitting package’s responsibility |
inst/stan/, src/
|
rstan::stan_model(),
cmdstanr::cmdstan_model() (user-managed) |
multisiteDGP installs without a C++ compiler; no compiled artifacts ship |
pp_check_rubin_normal() |
bayesplot::pp_check(),
posterior::summarise_draws()
|
Posterior diagnostics belong with the posterior, not the DGP |
loo_rubin_normal() |
loo::loo() on the fit object |
Model comparison is downstream of fitting |
The split makes both packages smaller. multisiteDGP is pure R and ships only the four generative layers plus diagnostics; whichever Bayesian engine fits best for a project lives in that engine’s package, not in a vendor lock-in wrapper (Walters, 2024).
4. MV1 — front door
The monolithic sim_multisite_data() call with many flat
arguments becomes either a preset piped into
sim_multisite() or a direct call with the same flat
arguments.
Before (siteBayes2):
library(siteBayes2)
df_old <- sim_multisite_data(
true_dist = "Mixture",
J = 50,
sigma_tau = 0.20,
delta = 5,
eps = 0.3,
ups = 2,
nj_mean = 40,
cv = 0.50,
nj_min = 5,
precision_dependence = FALSE,
set_seed = TRUE
)After (multisiteDGP — preset path):
dat <- preset_jebs_paper() |>
sim_multisite(seed = 123L)
print(dat, n = 3)
#> # A multisitedgp_data: 50 sites, paradigm = "site_size"
#> # Realized vs intended:
#> # I: realized=0.243 (no target)
#> # R: realized=17.000 (no target)
#> # sigma_tau: target=0.200, realized=0.199, PASS
#> # rho_S: target=0.000, realized=0.130, PASS
#> # rho_S_marg: realized=0.130 (no target)
#> # Feasibility: WARN (n_eff=12.853)
#> # A tibble: 50 × 7
#> site_index z_j tau_j tau_j_hat se_j se2_j n_j
#> <int> <dbl> <dbl> <dbl> <dbl> <dbl> <int>
#> 1 1 -1.19 -0.238 -0.0622 0.392 0.154 26
#> 2 2 1.10 0.219 -0.0135 0.603 0.364 11
#> 3 3 -0.504 -0.101 -0.886 0.5 0.25 16
#> # ℹ 47 more rows
#> # Use summary(df) for the full diagnostic report.The print method displays realized vs. intended diagnostics
(informativeness
,
heterogeneity ratio
,
residual heterogeneity
,
rank correlation
).
preset_jebs_paper()
is the JEBS replication preset; the sim_multisite()
front door applies it through the four layers.
5. MV2 — flat JEBS-style arguments
When the script needs the arguments visible (e.g. for a sweep), pass
them directly to sim_multisite().
The mixture parameters now live inside theta_G; everything
else is a named argument with the original meaning.
flat_dat <- sim_multisite(
J = 50L,
true_dist = "Mixture",
theta_G = list(delta = 5, eps = 0.3, ups = 2),
sigma_tau = 0.20,
nj_mean = 40,
cv = 0.50,
nj_min = 5L,
engine = "A1_legacy",
dependence = "none",
seed = 123L
)
head(tibble::as_tibble(flat_dat)[, c("site_index", "z_j", "tau_j", "tau_j_hat", "se2_j")], 3)
#> # A tibble: 3 × 5
#> site_index z_j tau_j tau_j_hat se2_j
#> <int> <dbl> <dbl> <dbl> <dbl>
#> 1 1 -1.19 -0.238 -0.0622 0.154
#> 2 2 1.10 0.219 -0.0135 0.364
#> 3 3 -0.504 -0.101 -0.886 0.25engine = "A1_legacy" reproduces the original
siteBayes2 site-size algorithm (truncated-and-rounded
gamma); the default engine = "A2_modern" uses
moment-matched truncated gamma — both hit the same
(nj_mean, cv) target, but the legacy engine matches JEBS
Table 3 row-for-row (Lee et al., 2025).
See M2: G-distribution
catalog for the
-shape
parameter convention theta_G enforces.
6. MV3 — covariate-aware and custom
gen_priorG() and gen_priorG2() collapse to
a single gen_effects(). The
covariate-aware path is an argument set (formula,
beta, data), not a separate function.
x <- data.frame(X1 = as.numeric(scale(seq_len(30L))))
effects <- gen_effects(
J = 30L,
true_dist = "Gaussian",
sigma_tau = 0.15,
formula = ~ X1,
beta = 0.30,
data = x
)
head(effects, 3)
#> # A tibble: 3 × 4
#> site_index z_j tau_j X1
#> <int> <dbl> <dbl> <dbl>
#> 1 1 -0.626 -0.588 -1.65
#> 2 2 0.184 -0.433 -1.53
#> 3 3 -0.836 -0.551 -1.42For a non-built-in
,
switch to true_dist = "User" and pass g_fn.
The callback returns standardized residuals by default; multisiteDGP
audits and rescales them.
my_g <- function(J, ...) {
z <- stats::rt(J, df = 5)
(z - mean(z)) / stats::sd(z)
}
custom_dat <- sim_multisite(
multisitedgp_design(
J = 40L,
true_dist = "User",
g_fn = my_g,
g_returns = "standardized"
),
seed = 44L
)
informativeness(custom_dat)
#> [1] 0.2984526The full
custom-
contract — including the g_returns = "raw" escape hatch —
is in M5: Custom G
distributions. The eight built-in shapes are catalogued in M2: G-distribution catalog,
which includes the gen_priorG rename map under each
shape.
7. MV4 — split dependence and observation
sim_observed_effects() used to combine dependence
alignment and observation sampling. multisiteDGP keeps those layers
separate so each can be inspected, customized, or audited.
effects <- gen_effects_gaussian(J = 20L, sigma_tau = 0.20)
margins <- gen_site_sizes(effects, J = 20L, nj_mean = 50, cv = 0.30)
dependent <- align_rank_corr(margins, rank_corr = 0.30, max_iter = 2000L)
observed <- gen_observations(dependent)
stats::cor(observed$z_j, observed$se2_j, method = "spearman")
#> [1] 0.3002258
head(observed, 3)
#> # A tibble: 3 × 7
#> site_index z_j tau_j n_j se2_j se_j tau_j_hat
#> <int> <dbl> <dbl> <int> <dbl> <dbl> <dbl>
#> 1 1 1.36 0.272 42 0.0952 0.309 0.418
#> 2 2 -0.103 -0.0206 54 0.0741 0.272 -0.214
#> 3 3 0.388 0.0775 58 0.0690 0.263 0.238Most users should still call sim_multisite();
the layered calls are useful when migrating custom internals or when
swapping a single layer (e.g. a copula dependence injection in place of
rank-corr alignment).
8. MV5 — Stan fitting moves downstream
multisiteDGP does not ship Stan models, Rcpp code, or Bayesian fitting wrappers. The package installs without a C++ compiler.
Before (siteBayes2):
library(siteBayes2)
fit_old <- fit_rubin_normal(
df = sites,
tau_hat_col = "tau_j_hat",
se2_col = "se2_j",
iter = 2000,
chains = 4
)After (multisiteDGP — adapter to metafor):
fit_dat <- sim_multisite(J = 30L, seed = 1L)
if (requireNamespace("metafor", quietly = TRUE)) {
meta_dat <- as_metafor(fit_dat)
fit_meta <- metafor::rma(yi = meta_dat$yi, vi = meta_dat$vi, method = "REML")
print(fit_meta)
} else {
message("Install metafor for the random-effects fit: install.packages('metafor')")
}
#>
#> Random-Effects Model (k = 30; tau^2 estimator: REML)
#>
#> tau^2 (estimated amount of total heterogeneity): 0.0419 (SE = 0.0334)
#> tau (square root of estimated tau^2 value): 0.2047
#> I^2 (total heterogeneity / total variability): 32.89%
#> H^2 (total variability / sampling variability): 1.49
#>
#> Test for Heterogeneity:
#> Q(df = 29) = 43.3550, p-val = 0.0422
#>
#> Model Results:
#>
#> estimate se zval pval ci.lb ci.ub
#> 0.0342 0.0662 0.5162 0.6057 -0.0956 0.1639
#>
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1The adapter renames the canonical columns to metafor’s
(yi, vi, sei) convention without touching the numerical
values. The full contract is in M6: Adapters and downstream
packages.
9. MV6 — baggr path
Bayesian random-effects fitting through baggr follows
the same pattern: adapt, then fit. include_truth = TRUE
keeps tau_j as tau_true so post-fit recovery
checks are one column away.
if (requireNamespace("baggr", quietly = TRUE)) {
bg <- as_baggr(fit_dat, include_truth = TRUE)
head(bg, 3)
} else {
message("Install baggr for the Bayesian random-effects fit: install.packages('baggr')")
}
#> # A tibble: 3 × 3
#> tau se tau_true
#> <dbl> <dbl> <dbl>
#> 1 0.923 0.436 -0.125
#> 2 0.0266 0.258 0.0367
#> 3 0.0320 0.289 -0.167as_baggr()
renames to (tau, se, tau_true). The fit call itself is
baggr::baggr(bg) — multisiteDGP does not wrap it.
10. MV7 — shrinkage helper
get_shrinkage_factor() is replaced by mean_shrinkage(),
now vectorized over nj_mean.
mean_shrinkage(
nj_mean = c(20, 50, 100),
sigma_tau = 0.15,
varY = 1,
p = 0.5,
R2 = 0.20
)
#> [1] 0.1232877 0.2601156 0.4128440The three values are the mean shrinkage factors at
.
Larger sites see stronger shrinkage toward the global mean. The
diagnostic also has a multisitedgp_data method that reads
varY and p from the design’s provenance.
11. MV8 — no legacy exports in multisiteDGP
multisiteDGP intentionally does not export siteBayes2
names. That keeps the new package small, pure-R, and DGP-only. If a
script needs backwards compatibility, the place for that is a thin
siteBayes2-compatibility patch package that imports
multisiteDGP and issues deprecation warnings — not multisiteDGP
itself.
legacy_names <- c(
"sim_multisite_data",
"gen_priorG",
"gen_priorG2",
"sim_sitesize_withinvar",
"sim_observed_effects",
"get_shrinkage_factor"
)
legacy_names %in% getNamespaceExports("multisiteDGP")
#> [1] FALSE FALSE FALSE FALSE FALSE FALSEAll six return FALSE, confirming the boundary.
12. Realized-design check on a migrated script
A migrated script should be checked the same way an originally-written multisiteDGP script is — by reading the realized-vs.-intended diagnostic block and the funnel plot. The two together confirm that the new pipeline reproduces the design intent of the old call.
migrated <- preset_jebs_paper() |> sim_multisite(seed = 123L)
summary(migrated)
#> multisiteDGP simulation diagnostics
#> ------------------------------------------------------------
#> A. Realized vs Intended
#> I (informativeness): 0.243 (target N/A) N/A [no target]
#> R (SE heterogeneity): 17.000 (target N/A) N/A [no target]
#> sigma_tau: 0.199 (target 0.200) PASS [rel=-0.4%]
#> GM(se^2): 0.125 (target N/A) N/A [no target]
#>
#> B. Dependence
#> rank_corr residual: 0.130 (target 0.000) PASS [delta=0.130]
#> rank_corr marginal: 0.130 (target N/A) N/A [residual target rows only; no finite target; status not assigned]
#> pearson_corr residual: 0.034 (target 0.000) WARN [delta=0.034]
#> pearson_corr marginal: 0.034 (target N/A) N/A [residual target rows only; no finite target; status not assigned]
#>
#> C. G shape fit
#> KS distance D_J: N/A (target 0.000) N/A [no target]
#> Bhattacharyya BC: N/A (target 1.000) N/A [no target]
#> Q-Q residual: N/A (target 0.000) N/A [order-statistic band metadata deferred]
#>
#> D. Operational feasibility
#> mean shrinkage S: 0.257 (target N/A) PASS [no target]
#> avg MOE (95%): 0.722 (target N/A) WARN [no target]
#> feasibility_index: 12.853 (target N/A) WARN [no target]
#> ------------------------------------------------------------
#> Overall: 3 PASS, 3 WARN, 0 FAIL.
#> Provenance: multisiteDGP 0.1.1 | paradigm=site_size | seed=123 | canonical_hash=7bb08e59be82c6e7 | design_hash=e3b4a0a694b82f67 | hash_algo=xxhash64 | R=4.6.0 | hooks=none
plot_funnel(migrated)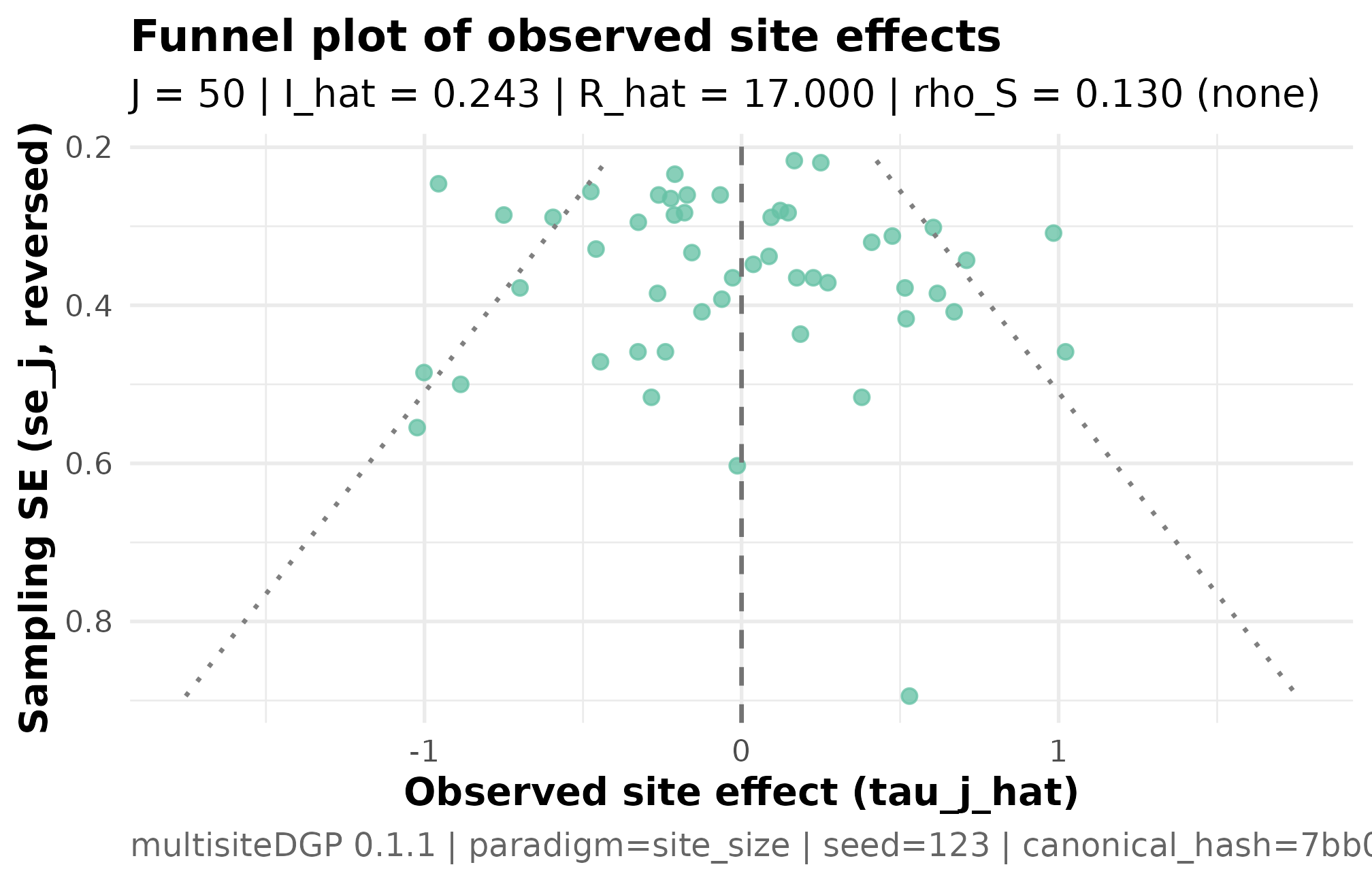
Funnel plot of the migrated JEBS-preset simulation. Each point is a
site; horizontal axis is the site SE, vertical axis is the observed
estimate tau_j_hat. The funnel widens with SE as expected
under the canonical factorization
.
No high-precision sites cluster at extreme estimates and no
low-precision sites cluster near zero, confirming the migrated pipeline
preserves the precision-effect independence target
.
The diagnostic block reports realized against the target (PASS) and realized against the target (PASS within tolerance). The funnel plot is the visual companion: symmetric scatter widening with SE confirms the precision-effect independence the JEBS preset targets.
13. Three points where the new API diverges, not just renames
Most patterns above are pure renames. Three are not.
-
Layer separation (MV4).
sim_observed_effects()mixed dependence injection and observation sampling. multisiteDGP separates them intoalign_rank_corr()/align_copula_corr()/align_hybrid_corr()(Layer 3) andgen_observations()(Layer 4). Three precision-injection methods replace the booleanprecision_dependence. The full contract — including which method hits the rank-correlation target most efficiently at given — is in M4: Precision dependence — three injection methods. -
Two front doors, two paradigms (MV2 and MV5).
siteBayes2treated direct-precision designs as a flag. multisiteDGP separates them into two front doors:sim_multisite()for the site-size-driven path (paradigm = "site_size"— sample sizes produce SEs ) andsim_meta()for the direct-precision path (paradigm = "direct"— informativeness and heterogeneity ratio produce SEs directly via a geometric grid). The construction-time check redirects mismatches at design time. See M3: Margin and SE models for the formal statement. -
g_fnstandardization audit (MV3).siteBayes2’s custom hook accepted any callback. multisiteDGP audits the callback’s output againstg_returns = "standardized"(default) or"raw"and rescales to the requestedsigma_tau; the audit can be turned off for development work viaaudit_g = FALSE. See M5: Custom G distributions.
14. Where to next
For an end-to-end migrated workflow:
- A1: Getting started — the basic four-step pipeline, the same pipeline a migrated script ends up running.
- A2: Choosing a preset — the eight presets cover the JEBS replication, two meta-analytic profiles, and small-area estimation; most migrated scripts can replace a flat argument list with a preset call.
For the methodological mapping behind the renames:
-
M2: G-distribution
catalog — the eight built-in
shapes and the
gen_priorG→gen_effectsparameter map per shape. -
M3: Margin and SE models —
the site-size-driven and direct-precision paths,
sim_multisite()vs.sim_meta(), and why the two are separated rather than flagged. -
M4: Precision
dependence — three injection methods — the rank, copula, and hybrid
alternatives that replace
precision_dependence = TRUE/FALSE.
References
The two-stage data-generating process and the canonical factorization that the renamed function set implements is from Lee et al. (2025). The empirical-Bayes shrinkage target the migrated adapter pipeline produces is the analysis goal in Walters (2024).
Acknowledgments
This research was supported by the Institute of Education Sciences, U.S. Department of Education, through Grant R305D240078 to the University of Alabama. The opinions expressed are those of the authors and do not represent views of the Institute or the U.S. Department of Education.
Session info
#> R version 4.6.0 (2026-04-24)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ggplot2_4.0.3 multisiteDGP_0.1.1
#>
#> loaded via a namespace (and not attached):
#> [1] gtable_0.3.6 xfun_0.57 bslib_0.10.0
#> [4] QuickJSR_1.9.2 ggrepel_0.9.8 inline_0.3.21
#> [7] lattice_0.22-9 mathjaxr_2.0-0 numDeriv_2016.8-1.1
#> [10] vctrs_0.7.3 tools_4.6.0 generics_0.1.4
#> [13] yulab.utils_0.2.4 parallel_4.6.0 stats4_4.6.0
#> [16] tibble_3.3.1 pkgconfig_2.0.3 Matrix_1.7-5
#> [19] checkmate_2.3.4 ggplotify_0.1.3 RColorBrewer_1.1-3
#> [22] S7_0.2.2 desc_1.4.3 RcppParallel_5.1.11-2
#> [25] lifecycle_1.0.5 compiler_4.6.0 farver_2.1.2
#> [28] textshaping_1.0.5 codetools_0.2-20 forestplot_3.2.0
#> [31] htmltools_0.5.9 sass_0.4.10 bayesplot_1.15.0
#> [34] yaml_2.3.12 pillar_1.11.1 pkgdown_2.2.0
#> [37] crayon_1.5.3 jquerylib_0.1.4 cachem_1.1.0
#> [40] StanHeaders_2.32.10 metadat_1.6-0 abind_1.4-8
#> [43] nlme_3.1-169 rstan_2.32.7 tidyselect_1.2.1
#> [46] digest_0.6.39 dplyr_1.2.1 labeling_0.4.3
#> [49] fastmap_1.2.0 grid_4.6.0 cli_3.6.6
#> [52] metafor_5.0-1 magrittr_2.0.5 loo_2.9.0
#> [55] pkgbuild_1.4.8 withr_3.0.2 scales_1.4.0
#> [58] backports_1.5.1 rappdirs_0.3.4 rmarkdown_2.31
#> [61] matrixStats_1.5.0 gridExtra_2.3 ragg_1.5.2
#> [64] evaluate_1.0.5 knitr_1.51 baggr_0.8
#> [67] nleqslv_3.3.7 gridGraphics_0.5-1 rstantools_2.6.0
#> [70] rlang_1.2.0 Rcpp_1.1.1-1.1 glue_1.8.1
#> [73] jsonlite_2.0.0 R6_2.6.1 systemfonts_1.3.2
#> [76] fs_2.1.0