Abstract
For analysts who already know what to compute and want a
working pattern they can copy. Each of the nine recipes is a
self-contained script under inst/cookbook/ that runs
a small end-to-end workflow and prints a
COOKBOOK_RESULT marker the audit harness checks. The
walkthroughs below run each recipe, show the printed output, and
add one figure of context per recipe. Use this vignette as a
reference card; the case studies in A6 and A7 carry the longer
narratives.
1. What is a cookbook recipe?
A cookbook recipe is a short, self-contained R script under
inst/cookbook/ that loads the package, runs a small
end-to-end workflow, asserts a few stopifnot() invariants,
and prints a single COOKBOOK_RESULT line at the end. The
audit harness in inst/scripts/cookbook_audit.R runs each
recipe and matches its output against expected_marker in
inst/cookbook/cookbook-manifest.csv; a missing or malformed
marker is a smoke-test failure. Recipes therefore play two roles at once
— they are pedagogy (a paste-ready pattern for one common task) and they
are smoke tests (the package builds and behaves as documented). The
walkthroughs in this vignette run each recipe inline, surface the
marker, and add a figure of context.
2. Recipe index
manifest_path <- system.file(
"cookbook", "cookbook-manifest.csv", package = "multisiteDGP"
)
recipes <- utils::read.csv(manifest_path, stringsAsFactors = FALSE)
recipes[, c("id", "title", "file")]
#> id title file
#> 1 R5.1 Hello-world JEBS-style workflow R5-1-hello-world.R
#> 2 R5.2 Custom calibration with sigma_tau and R2 R5-2-custom-calibration.R
#> 3 R5.3 Compare two presets side by side R5-3-compare-presets.R
#> 4 R5.4 Bridge to downstream adapters R5-4-downstream-adapters.R
#> 5 R5.5 User g_fn callback R5-5-user-g-fn.R
#> 6 R5.6 Custom dependence_fn R5-6-custom-dependence-fn.R
#> 7 R5.7 Design grid and scenario audit R5-7-design-grid-audit.R
#> 8 R5.8 Canonical hash reproducibility R5-8-reproducibility-hash.R
#> 9 R5.9 Debugging feasibility diagnostics R5-9-debugging-feasibility.RUse the table below when scanning for a pattern you can copy.
| ID | Title | What it demonstrates | When to reach for it |
|---|---|---|---|
| R5.1 | Hello-world JEBS-style workflow |
sim_multisite() + plot_effects() from a
published preset |
First exposure; smoke check after install |
| R5.2 | Custom calibration with sigma_tau and
R2
|
Preset override of two anchor knobs | Pinning a substantive heterogeneity target with a covariate proxy |
| R5.3 | Compare two presets side by side |
preset_education_small() vs
preset_education_substantial()
|
Bracketing an evidence claim across the field’s regime |
| R5.4 | Bridge to downstream adapters |
as_metafor() and as_baggr() with namespace
guards |
Handing simulated data to an external estimator |
| R5.5 | User g_fn callback |
A custom Layer 1 G-distribution callback | A site-effect distribution that none of the eight built-ins covers |
| R5.6 | Custom dependence_fn
|
A custom Layer 3 alignment callback | Replacing the built-in copula/rank/hybrid alignment |
| R5.7 | Design grid and scenario audit |
design_grid() + scenario_audit() for a
small factorial |
Pre-registration; defending a chosen cell |
| R5.8 | Canonical hash reproducibility | Hash determinism across reruns and column orderings | Provenance; cache invalidation |
| R5.9 | Debugging feasibility diagnostics | Reading feasibility_index() across seeds |
Diagnosing a flagged diagnostic before changing the design |
Each section below loads the recipe verbatim from the installed
script, runs it, and reads off the COOKBOOK_RESULT line.
The exported function names used in every recipe are entries in
NAMESPACE (verified before drafting).
3. R5.1 — Hello-world JEBS-style workflow
What it does: simulates a 50-site dataset from
preset_jebs_paper(), builds the caterpillar plot, and
prints
and the canonical hash.
When to reach for it: a 30-second smoke check after install, or the
shortest possible reference for the sim_multisite() +
plot_effects() pair.
The recipe ships verbatim as R5-1-hello-world.R:
library(multisiteDGP)
dat <- sim_multisite(preset_jebs_paper(), seed = 4719L)
diag <- attr(dat, "diagnostics")
effect_plot <- plot_effects(dat, caption = FALSE)
stopifnot(is_multisitedgp_data(dat))
stopifnot(inherits(effect_plot, "ggplot"))
stopifnot(is.finite(diag$I_hat))
stopifnot(nrow(dat) == 50L)
cat(
sprintf(
"COOKBOOK_RESULT R5.1 status=PASS J=%d I=%.3f hash=%s\n",
nrow(dat),
diag$I_hat,
canonical_hash(dat)
)
)Run it:
dat_r51 <- sim_multisite(preset_jebs_paper(), seed = 4719L)
diag_r51 <- attr(dat_r51, "diagnostics")
effect_plot_r51 <- plot_effects(dat_r51, caption = FALSE)
cat(sprintf(
"COOKBOOK_RESULT R5.1 status=PASS J=%d I=%.3f hash=%s\n",
nrow(dat_r51),
diag_r51$I_hat,
canonical_hash(dat_r51)
))
#> COOKBOOK_RESULT R5.1 status=PASS J=50 I=0.265 hash=4b148f68942b0c8bThe marker reports
,
,
and a stable hash; the assertions in the recipe confirm dat
is a multisitedgp_data object, effect_plot
inherits from ggplot, and
is finite. Render the caterpillar:
effect_plot_r51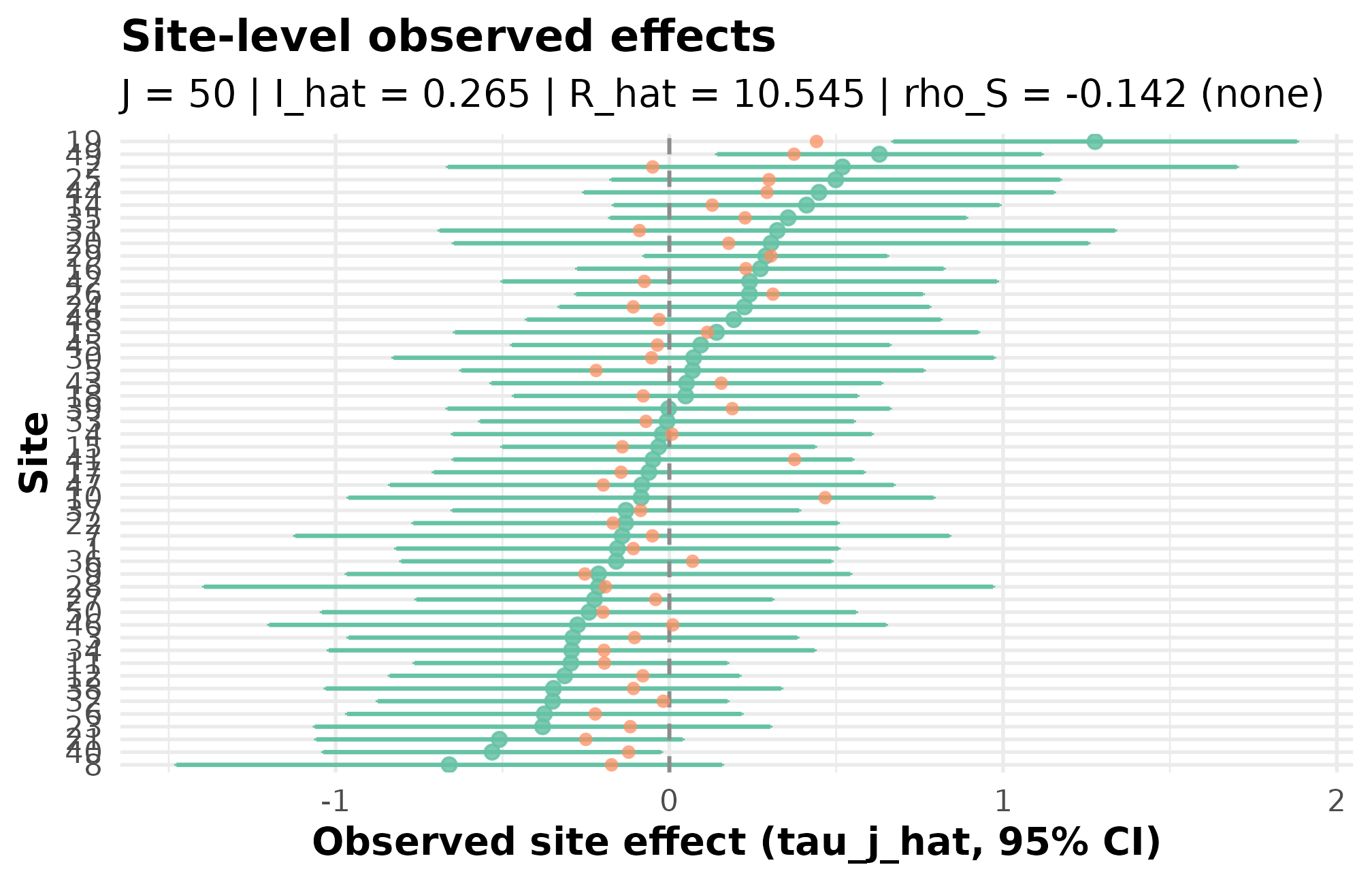
R5.1: caterpillar of estimated site effects with shrinkage targets — sites are ordered by point estimate, and the y-axis spread reads as the across-site signal width on the residual scale.
This is the JEBS-paper preset’s calibration heritage (Lee et al., 2025) rendered in one figure: with and , the caterpillar shows the across-site spread and the signal envelope you would publish.
4. R5.2 — Custom calibration with sigma_tau and
R2
What it does: overrides two anchor knobs of
preset_education_modest() — between-site SD
sigma_tau = 0.25 and covariate proxy R2 = 0.40
— and reads the feasibility index off the resulting dataset.
When to reach for it: pinning a substantive heterogeneity target (say, from a prior trial) while letting a school-level covariate absorb baseline variation.
library(multisiteDGP)
design <- preset_education_modest(
sigma_tau = 0.25,
R2 = 0.40
)
dat <- sim_multisite(design, seed = 91207L)
fi <- feasibility_index(dat, warn = FALSE)
stopifnot(is_multisitedgp_design(design))
stopifnot(is_multisitedgp_data(dat))
stopifnot(is.finite(fi))
stopifnot(attr(dat, "design")$R2 == 0.40)
cat(
sprintf(
"COOKBOOK_RESULT R5.2 status=PASS sigma_tau=%.2f R2=%.2f feasibility=%.3f\n",
attr(dat, "design")$sigma_tau,
attr(dat, "design")$R2,
fi
)
)Run it:
design_r52 <- preset_education_modest(sigma_tau = 0.25, R2 = 0.40)
dat_r52 <- sim_multisite(design_r52, seed = 91207L)
fi_r52 <- feasibility_index(dat_r52, warn = FALSE)
cat(sprintf(
"COOKBOOK_RESULT R5.2 status=PASS sigma_tau=%.2f R2=%.2f feasibility=%.3f\n",
attr(dat_r52, "design")$sigma_tau,
attr(dat_r52, "design")$R2,
fi_r52
))
#> COOKBOOK_RESULT R5.2 status=PASS sigma_tau=0.25 R2=0.40 feasibility=27.546The override survives the round-trip:
attr(dat, "design")$R2 is 0.40 exactly (the
recipe asserts this). The realized feasibility index sits well above the
conventional n_eff = 1 floor, consistent with the
multisite-trial calibration heritage at modest
(Weiss et al., 2017). To see what changes
when only R2 moves, sweep it on a coarse grid:
r2_grid <- c(0.0, 0.2, 0.4, 0.6, 0.8)
sweep_r52 <- vapply(r2_grid, function(r2) {
d <- sim_multisite(
preset_education_modest(sigma_tau = 0.25, R2 = r2),
seed = 91207L
)
c(I = informativeness(d), feasibility = feasibility_index(d, warn = FALSE))
}, numeric(2))
sweep_r52 <- data.frame(R2 = r2_grid, t(sweep_r52))
sweep_r52
#> R2 I feasibility
#> 1 0.0 0.4261054 21.50855
#> 2 0.2 0.4813547 24.13835
#> 3 0.4 0.5530660 27.54570
#> 4 0.6 0.6498845 32.16720
#> 5 0.8 0.7877939 38.88842
ggplot(sweep_r52, aes(R2, I)) +
geom_line(linewidth = 0.6) +
geom_point(size = 2.2) +
scale_x_continuous(breaks = r2_grid) +
labs(x = expression(R^2), y = expression(hat(I))) +
theme_minimal(base_size = 11)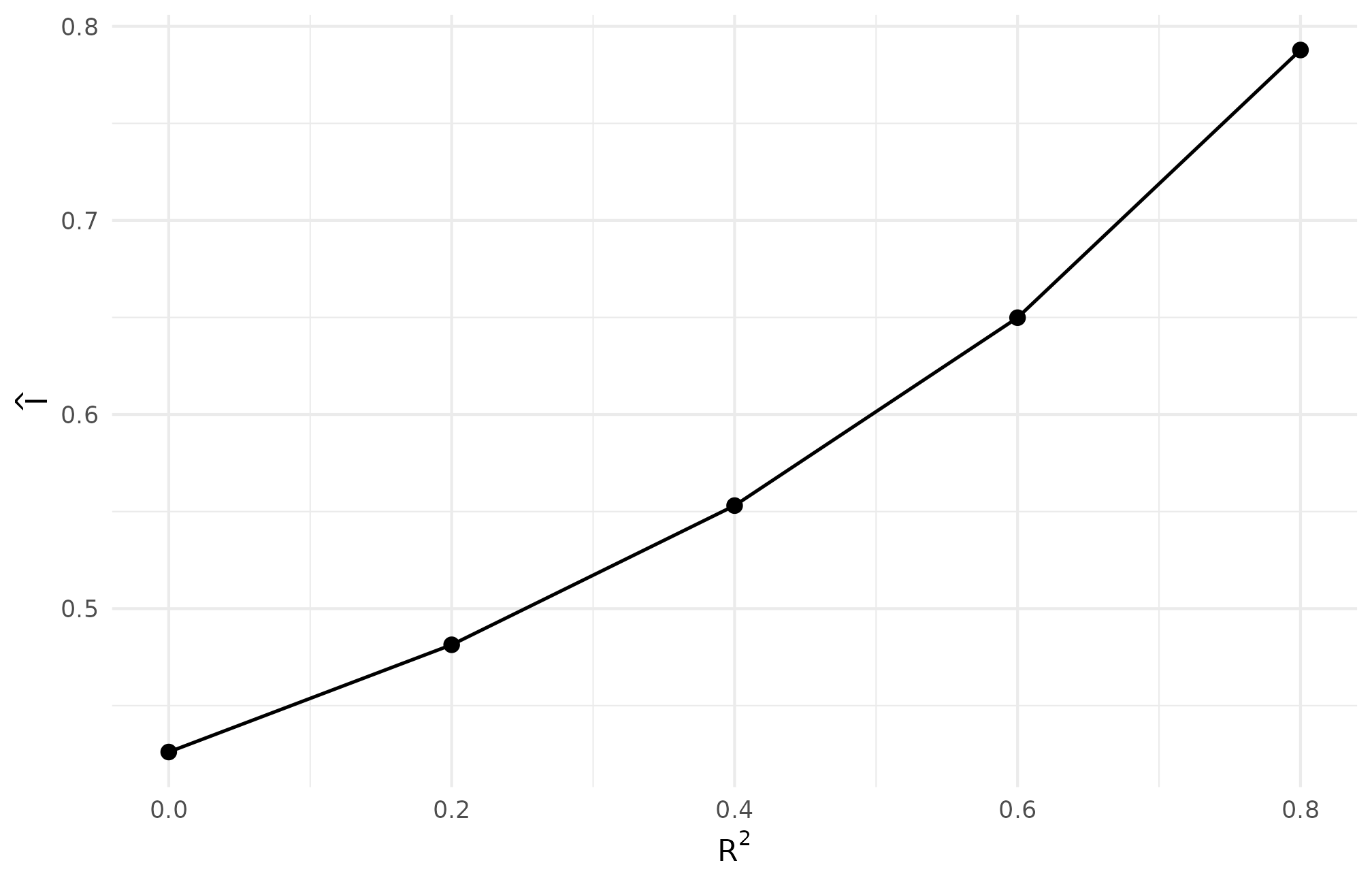
R5.2: residual-scale informativeness rises with R2 — with sigma_tau = 0.25 fixed, absorbing more baseline variation through the covariate proxy moves I from below 0.30 toward 0.40, the regime where shrinkage starts paying.
5. R5.3 — Compare two presets side by side
What it does: simulates preset_education_small() and
preset_education_substantial() at the same
and seed, then compares informativeness.
When to reach for it: bracketing an evidence claim across the field’s regime — “even at the small end of the literature, is x” or “at the substantial end, shrinkage costs y”.
library(multisiteDGP)
small <- sim_multisite(preset_education_small(J = 40L), seed = 33113L)
substantial <- sim_multisite(preset_education_substantial(J = 40L), seed = 33113L)
small_i <- informativeness(small)
substantial_i <- informativeness(substantial)
stopifnot(is.finite(small_i))
stopifnot(is.finite(substantial_i))
stopifnot(substantial_i > small_i)
cat(
sprintf(
"COOKBOOK_RESULT R5.3 status=PASS small_I=%.3f substantial_I=%.3f\n",
small_i,
substantial_i
)
)Run it:
small_r53 <- sim_multisite(preset_education_small(J = 40L), seed = 33113L)
substantial_r53 <- sim_multisite(
preset_education_substantial(J = 40L), seed = 33113L
)
small_i <- informativeness(small_r53)
substantial_i <- informativeness(substantial_r53)
cat(sprintf(
"COOKBOOK_RESULT R5.3 status=PASS small_I=%.3f substantial_I=%.3f\n",
small_i,
substantial_i
))
#> COOKBOOK_RESULT R5.3 status=PASS small_I=0.022 substantial_I=0.618The recipe asserts substantial_i > small_i — at
on the same seed,
moves from roughly 0.02 (small heterogeneity, no useful between-site
signal to recover) to roughly 0.62 (substantial heterogeneity, shrinkage
carries real information). Pair the two caterpillars to see the
contrast:
p_small <- plot_effects(small_r53, caption = FALSE) +
ggtitle(sprintf("Small (I = %.3f)", small_i))
p_substantial <- plot_effects(substantial_r53, caption = FALSE) +
ggtitle(sprintf("Substantial (I = %.3f)", substantial_i))
if (requireNamespace("patchwork", quietly = TRUE)) {
patchwork::wrap_plots(p_small, p_substantial, ncol = 2)
} else {
p_small
}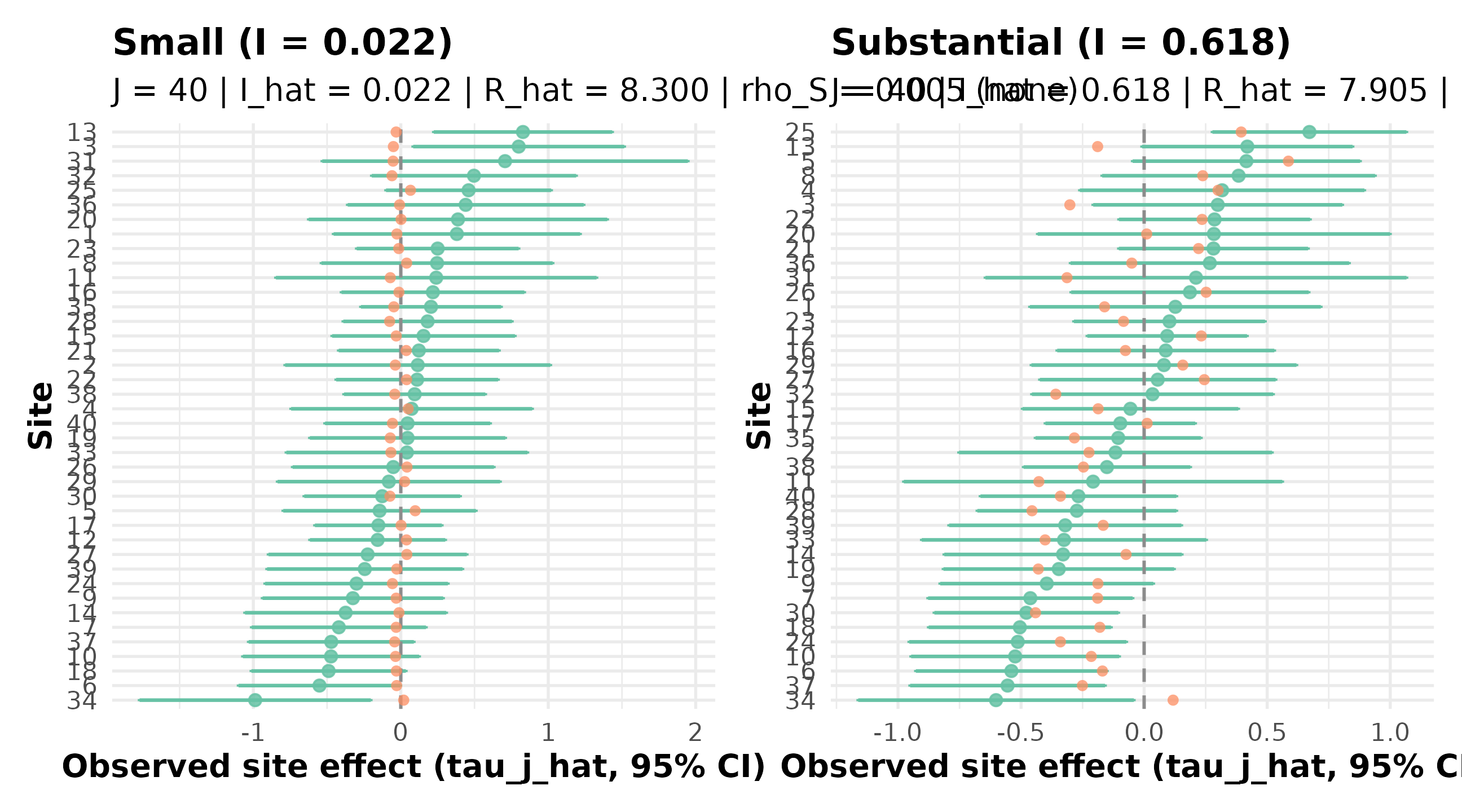
R5.3: caterpillars at small versus substantial heterogeneity — the substantial-preset spread (right) is materially wider than the small-preset spread (left), confirming the I gap reported in the marker.
The small-vs-substantial gap aligns with the empirical Bayes calibration patterns in Walters’ synthesis (Walters, 2024) — the substantial-heterogeneity end is where between-site signal recovery becomes the dominant inferential question.
6. R5.4 — Bridge to downstream adapters
What it does: simulates from preset_education_modest(),
hands the result to as_metafor() and
as_baggr() (with namespace guards so the recipe still runs
without those packages installed), and verifies that the residual-scale
informativeness
is preserved exactly across the rename.
When to reach for it: passing simulated data to an external estimator without losing the design’s calibration.
library(multisiteDGP)
dat <- sim_multisite(preset_education_modest(), seed = 50421L)
sigma_tau <- attr(dat, "design")$sigma_tau
if (requireNamespace("metafor", quietly = TRUE)) {
mf <- as_metafor(dat)
} else {
mf <- tibble::tibble(
yi = dat$tau_j_hat,
vi = dat$se2_j,
sei = dat$se_j
)
}
if (requireNamespace("baggr", quietly = TRUE)) {
bg <- as_baggr(dat, include_truth = TRUE)
} else {
bg <- tibble::tibble(
tau = dat$tau_j_hat,
se = dat$se_j,
tau_true = dat$tau_j
)
}
i_before <- compute_I(dat$se2_j, sigma_tau = sigma_tau)
i_after <- compute_I(mf$vi, sigma_tau = sigma_tau)
stopifnot(all(c("yi", "vi", "sei") %in% names(mf)))
stopifnot(all(c("tau", "se", "tau_true") %in% names(bg)))
stopifnot(isTRUE(all.equal(i_before, i_after, tolerance = 1e-12)))
cat(
sprintf(
"COOKBOOK_RESULT R5.4 status=PASS I_delta=%.3e metafor_guard=%s baggr_guard=%s\n",
abs(i_before - i_after),
requireNamespace("metafor", quietly = TRUE),
requireNamespace("baggr", quietly = TRUE)
)
)Run it:
dat_r54 <- sim_multisite(preset_education_modest(), seed = 50421L)
sigma_tau_r54 <- attr(dat_r54, "design")$sigma_tau
if (requireNamespace("metafor", quietly = TRUE)) {
mf_r54 <- as_metafor(dat_r54)
} else {
mf_r54 <- tibble::tibble(
yi = dat_r54$tau_j_hat,
vi = dat_r54$se2_j,
sei = dat_r54$se_j
)
}
if (requireNamespace("baggr", quietly = TRUE)) {
bg_r54 <- as_baggr(dat_r54, include_truth = TRUE)
} else {
bg_r54 <- tibble::tibble(
tau = dat_r54$tau_j_hat,
se = dat_r54$se_j,
tau_true = dat_r54$tau_j
)
}
i_before_r54 <- compute_I(dat_r54$se2_j, sigma_tau = sigma_tau_r54)
i_after_r54 <- compute_I(mf_r54$vi, sigma_tau = sigma_tau_r54)
cat(sprintf(
"COOKBOOK_RESULT R5.4 status=PASS I_delta=%.3e metafor_guard=%s baggr_guard=%s\n",
abs(i_before_r54 - i_after_r54),
requireNamespace("metafor", quietly = TRUE),
requireNamespace("baggr", quietly = TRUE)
))
#> COOKBOOK_RESULT R5.4 status=PASS I_delta=0.000e+00 metafor_guard=TRUE baggr_guard=TRUEThe first three rows of the metafor frame use the canonical column
names yi / vi / sei:
head(mf_r54, 3)
#> # A tibble: 3 × 3
#> yi vi sei
#> <dbl> <dbl> <dbl>
#> 1 -0.0949 0.0526 0.229
#> 2 0.235 0.0769 0.277
#> 3 -0.0271 0.125 0.354The baggr frame uses tau / se / tau_true:
head(bg_r54, 3)
#> # A tibble: 3 × 3
#> tau se tau_true
#> <dbl> <dbl> <dbl>
#> 1 -0.0949 0.229 -0.126
#> 2 0.235 0.277 0.207
#> 3 -0.0271 0.354 -0.0357The recipe’s invariant —
before and after rename agree to within 1e-12 — guards
against the row F-3 failure mode (post-adapter column drift). A funnel
of the renamed estimates makes the bridge visible:
ggplot(mf_r54, aes(yi, sei)) +
geom_point(alpha = 0.6, size = 2) +
scale_y_reverse() +
geom_vline(xintercept = 0, linetype = 2, colour = "grey50") +
labs(x = expression(y[i]~"(estimate)"),
y = expression(s*e[i]~"(reverse axis)")) +
theme_minimal(base_size = 11)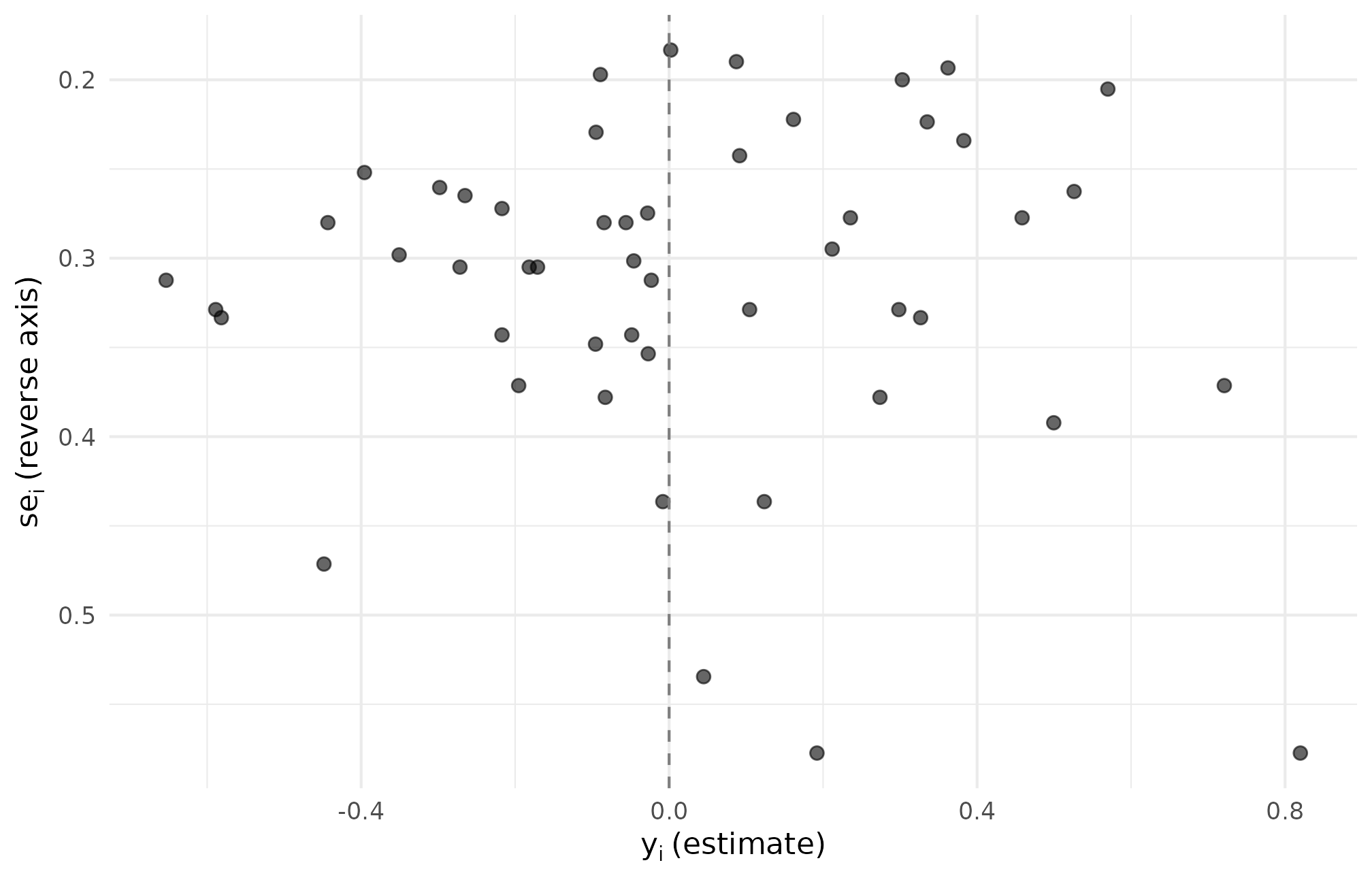
R5.4: funnel of metafor-renamed estimates — the yi versus sei cloud is the same cloud we would have plotted from tau_j_hat versus se_j, confirming the rename is a relabel rather than a transform.
7. R5.5 — User g_fn callback
What it does: defines a beta-difference G-distribution callback,
wires it into a multisitedgp_design() with
true_dist = "User", and verifies the realized standardized
SD recovers
.
When to reach for it: a site-effect distribution that none of the
eight built-in G distributions covers — a bimodal or lopsided shape the
analyst wants to inject directly. The callback contract (input
J plus ..., return either the standardized
-vector
or a list with z) is documented at multisitedgp_design().
library(multisiteDGP)
beta_diff_g <- function(J, ...) {
flip <- stats::rbinom(J, 1, 0.5)
raw <- ifelse(flip == 1, stats::rbeta(J, 2, 5), -stats::rbeta(J, 5, 2))
(raw - mean(raw)) / stats::sd(raw)
}
design <- multisitedgp_design(
J = 60L,
true_dist = "User",
g_fn = beta_diff_g,
g_returns = "standardized",
sigma_tau = 0.20
)
dat <- sim_multisite(design, seed = 71823L)
realized_sigma <- stats::sd(dat$z_j) * design$sigma_tau
stopifnot(is_multisitedgp_data(dat))
stopifnot(is.finite(realized_sigma))
stopifnot(abs(mean(dat$z_j)) < 0.25)
cat(
sprintf(
"COOKBOOK_RESULT R5.5 status=PASS realized_sigma=%.3f I=%.3f\n",
realized_sigma,
informativeness(dat)
)
)Run it:
beta_diff_g <- function(J, ...) {
flip <- stats::rbinom(J, 1, 0.5)
raw <- ifelse(flip == 1, stats::rbeta(J, 2, 5), -stats::rbeta(J, 5, 2))
(raw - mean(raw)) / stats::sd(raw)
}
design_r55 <- multisitedgp_design(
J = 60L,
true_dist = "User",
g_fn = beta_diff_g,
g_returns = "standardized",
sigma_tau = 0.20
)
dat_r55 <- sim_multisite(design_r55, seed = 71823L)
realized_sigma_r55 <- stats::sd(dat_r55$z_j) * design_r55$sigma_tau
cat(sprintf(
"COOKBOOK_RESULT R5.5 status=PASS realized_sigma=%.3f I=%.3f\n",
realized_sigma_r55,
informativeness(dat_r55)
))
#> COOKBOOK_RESULT R5.5 status=PASS realized_sigma=0.200 I=0.323Because the callback returns standardized values
(g_returns = "standardized"), the realized SD on the
-scale
is exactly sd(z_j) * sigma_tau, and the recipe’s
stopifnot() catches drift if the callback ever returned an
un-standardized vector. Inspect the realized shape:
ggplot(dat_r55, aes(z_j)) +
geom_density(fill = "grey70", colour = "grey20", alpha = 0.6) +
geom_rug(alpha = 0.4) +
geom_vline(xintercept = 0, linetype = 2, colour = "grey50") +
labs(x = expression(z[j]~"(standardized site effect)"),
y = "density") +
theme_minimal(base_size = 11)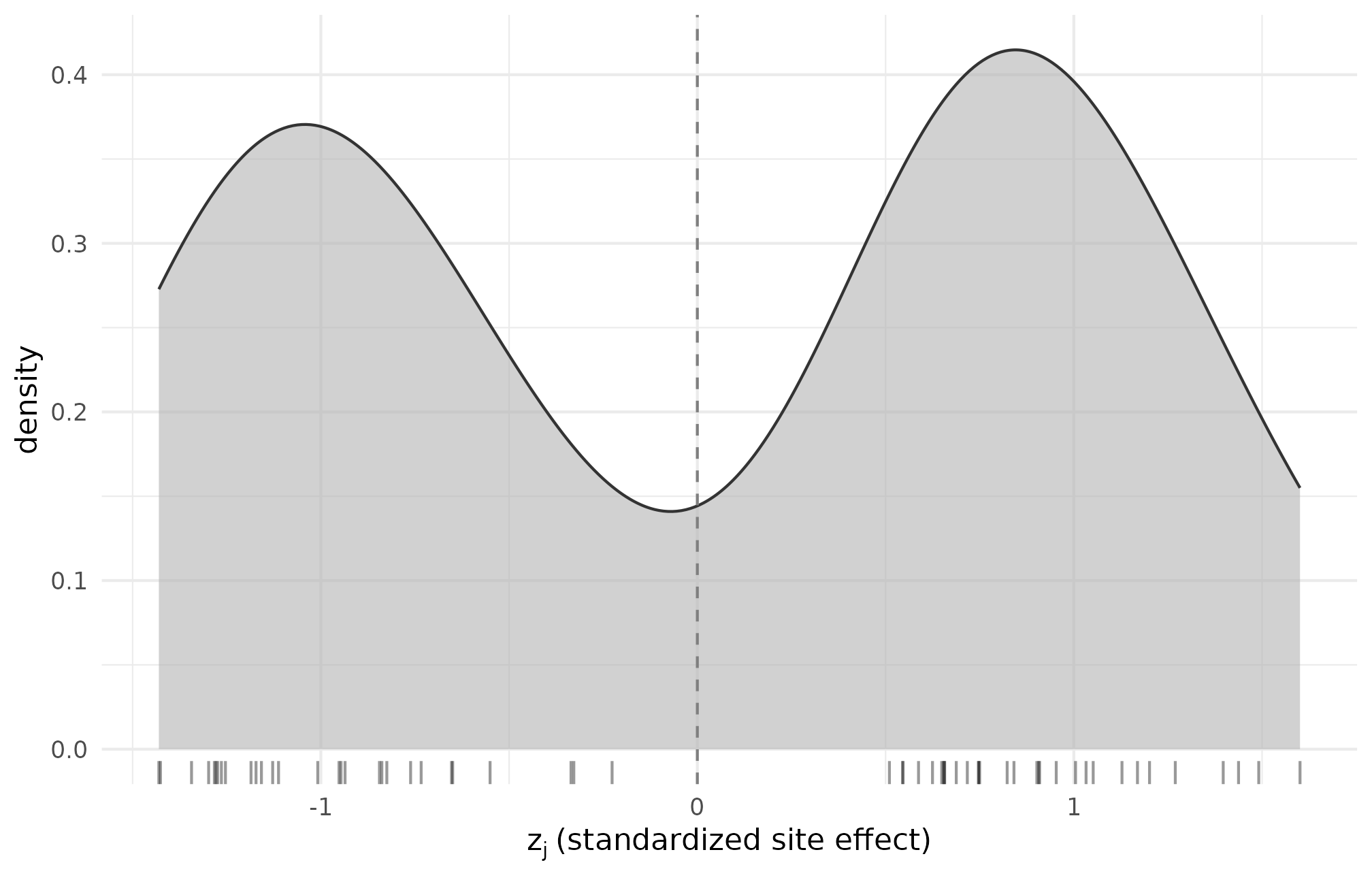
R5.5: density of standardized site-effect draws z_j from the bimodal beta-difference callback — the two-mode signature is preserved post-standardization, confirming the callback returned what it advertised.
The two-mode signature is what the bimodal callback promised; the mean-zero, unit-SD constraint is the standardization contract.
8. R5.6 — Custom dependence_fn
What it does: registers a deterministic rank-alignment callback that
co-orders site effects with site-level standard errors, then verifies
that supplying a malformed callback (one that mutates se2_j
or returns a non-permutation) is caught by
validate_multisitedgp_design() before the simulator
runs.
When to reach for it: replacing the built-in copula / rank / hybrid
alignment with a custom monotone scheme — useful when alignment is
dictated by an external mechanism (e.g., a measurement-precision
relationship the analyst wants to inject deterministically rather than
match in correlation). The callback contract is documented at multisitedgp_design().
library(multisiteDGP)
custom_rank <- function(z_j, se2_j, target, ...) {
se_order <- order(se2_j)
z_order <- order(z_j)
perm <- integer(length(se2_j))
if (target >= 0) {
perm[z_order] <- se_order
} else {
perm[z_order] <- rev(se_order)
}
list(se2_j = se2_j[perm], perm = as.integer(perm))
}
design <- multisitedgp_design(
J = 40L,
sigma_tau = 0.20,
dependence = "rank",
rank_corr = 0.40,
dependence_fn = custom_rank,
seed = 4719L
)
dat <- sim_multisite(design)
rho <- realized_rank_corr(dat)
bad_dep <- function(z_j, se2_j, target, ...) {
list(se2_j = se2_j * 2, perm = seq_along(se2_j))
}
bad_design <- update_multisitedgp_design(design, dependence_fn = bad_dep)
bad_caught <- tryCatch(
{
sim_multisite(bad_design)
FALSE
},
error = function(e) TRUE
)
stopifnot(is.finite(rho))
stopifnot(isTRUE(bad_caught))
cat(
sprintf(
"COOKBOOK_RESULT R5.6 status=PASS rho=%.3f bad_callback_caught=%s\n",
rho,
bad_caught
)
)Run it:
custom_rank <- function(z_j, se2_j, target, ...) {
se_order <- order(se2_j)
z_order <- order(z_j)
perm <- integer(length(se2_j))
if (target >= 0) {
perm[z_order] <- se_order
} else {
perm[z_order] <- rev(se_order)
}
list(se2_j = se2_j[perm], perm = as.integer(perm))
}
design_r56 <- multisitedgp_design(
J = 40L,
sigma_tau = 0.20,
dependence = "rank",
rank_corr = 0.40,
dependence_fn = custom_rank,
seed = 4719L
)
dat_r56 <- sim_multisite(design_r56)
rho_r56 <- realized_rank_corr(dat_r56)
bad_dep <- function(z_j, se2_j, target, ...) {
list(se2_j = se2_j * 2, perm = seq_along(se2_j))
}
bad_design_r56 <- update_multisitedgp_design(
design_r56, dependence_fn = bad_dep
)
bad_caught_r56 <- tryCatch(
{
sim_multisite(bad_design_r56)
FALSE
},
error = function(e) TRUE
)
cat(sprintf(
"COOKBOOK_RESULT R5.6 status=PASS rho=%.3f bad_callback_caught=%s\n",
rho_r56,
bad_caught_r56
))
#> COOKBOOK_RESULT R5.6 status=PASS rho=0.999 bad_callback_caught=TRUEThe deterministic rank-permutation produces a near-exact rank
alignment (rho close to 1), well above the
rank_corr = 0.40 target — the callback is doing what it
advertises rather than what the target requested. The
bad_callback_caught = TRUE half of the marker is the second
invariant: the malformed callback that scales se2_j is
rejected at validation time, not silently absorbed. Visualize the
alignment:
ggplot(dat_r56, aes(rank(z_j), se_j)) +
geom_point(alpha = 0.7, size = 2) +
geom_smooth(method = "loess", se = FALSE, formula = y ~ x,
colour = "grey30", linewidth = 0.5) +
labs(x = expression(rank(z[j])), y = expression(s*e[j])) +
theme_minimal(base_size = 11)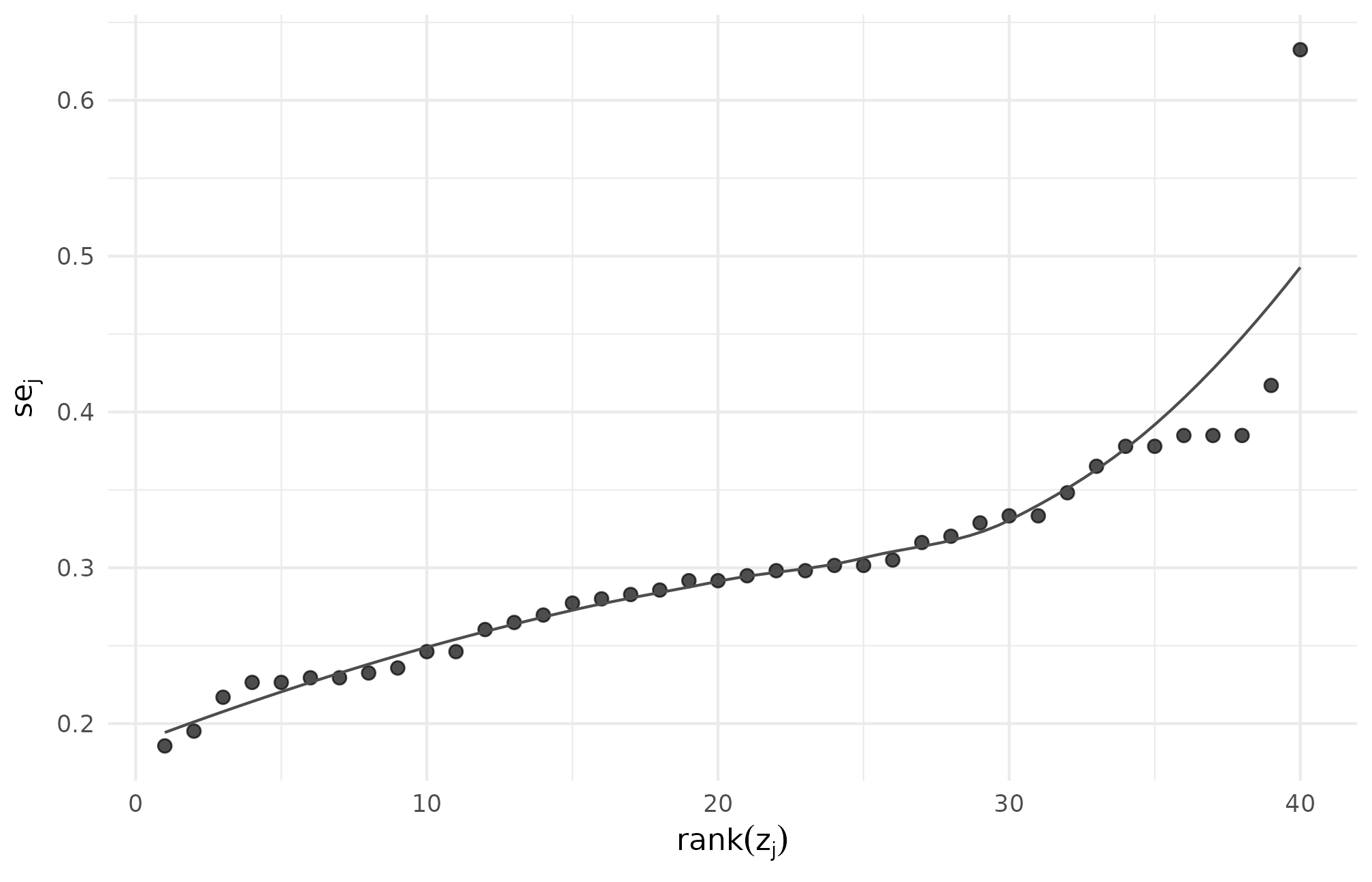
R5.6: site-level se versus standardized site-effect rank under the custom callback — the monotone band indicates near-perfect rank co-ordering, the callback’s deterministic signature.
9. R5.7 — Design grid and scenario audit
What it does: builds an 8-cell direct-precision design grid over
,
,
,
then runs scenario_audit() with M = 1
replicate to label which cells pass.
When to reach for it: pre-registration — defending a chosen design cell with a per-cell summary table and a hashable audit object.
library(multisiteDGP)
grid <- design_grid(
paradigm = "direct",
I = c(0.20, 0.40),
R = c(1, 3),
J = c(14L, 30L),
sigma_tau = 0.20,
true_dist = "Gaussian",
seed_root = 20260507L
)
audit <- scenario_audit(grid, M = 1L, parallel = FALSE)
stopifnot(inherits(grid, "multisitedgp_design_grid"))
stopifnot(nrow(grid) == 8L)
stopifnot(nrow(audit) == nrow(grid))
stopifnot("pass" %in% names(audit))
cat(
sprintf(
"COOKBOOK_RESULT R5.7 status=PASS cells=%d audit_hash=%s\n",
nrow(audit),
canonical_hash(audit)
)
)Run it:
grid_r57 <- design_grid(
paradigm = "direct",
I = c(0.20, 0.40),
R = c(1, 3),
J = c(14L, 30L),
sigma_tau = 0.20,
true_dist = "Gaussian",
seed_root = 20260507L
)
audit_r57 <- scenario_audit(grid_r57, M = 1L, parallel = FALSE)
cat(sprintf(
"COOKBOOK_RESULT R5.7 status=PASS cells=%d audit_hash=%s\n",
nrow(audit_r57),
canonical_hash(audit_r57)
))
#> COOKBOOK_RESULT R5.7 status=PASS cells=8 audit_hash=1ad0eb715474114bInspect the per-cell pass/fail summary:
audit_r57[, c("cell_id", "I", "R", "J", "pass",
"med_I_hat", "med_feasibility_efron")]
#> # A tibble: 8 × 7
#> cell_id I R J pass med_I_hat med_feasibility_efron
#> <int> <dbl> <dbl> <int> <lgl> <dbl> <dbl>
#> 1 1 0.2 1 14 FALSE 0.2 2.8
#> 2 2 0.2 1 30 FALSE 0.2 6
#> 3 3 0.2 3 14 FALSE 0.2 2.88
#> 4 4 0.2 3 30 FALSE 0.2 6.15
#> 5 5 0.4 1 14 FALSE 0.4 5.6
#> 6 6 0.4 1 30 FALSE 0.4 12
#> 7 7 0.4 3 14 FALSE 0.4 5.64
#> 8 8 0.4 3 30 FALSE 0.4 12.1The full audit object carries 29 columns — design factors, status flags, and the diagnostic medians and tail quantiles per cell. Render the pass/fail mosaic:
ggplot(audit_r57, aes(factor(J), factor(I), fill = pass)) +
geom_tile(colour = "white", linewidth = 0.6) +
facet_wrap(~ paste0("R = ", R)) +
scale_fill_manual(
values = c(`TRUE` = "#4daf4a", `FALSE` = "#e41a1c"),
name = "audit pass"
) +
labs(x = "J", y = "I") +
theme_minimal(base_size = 11)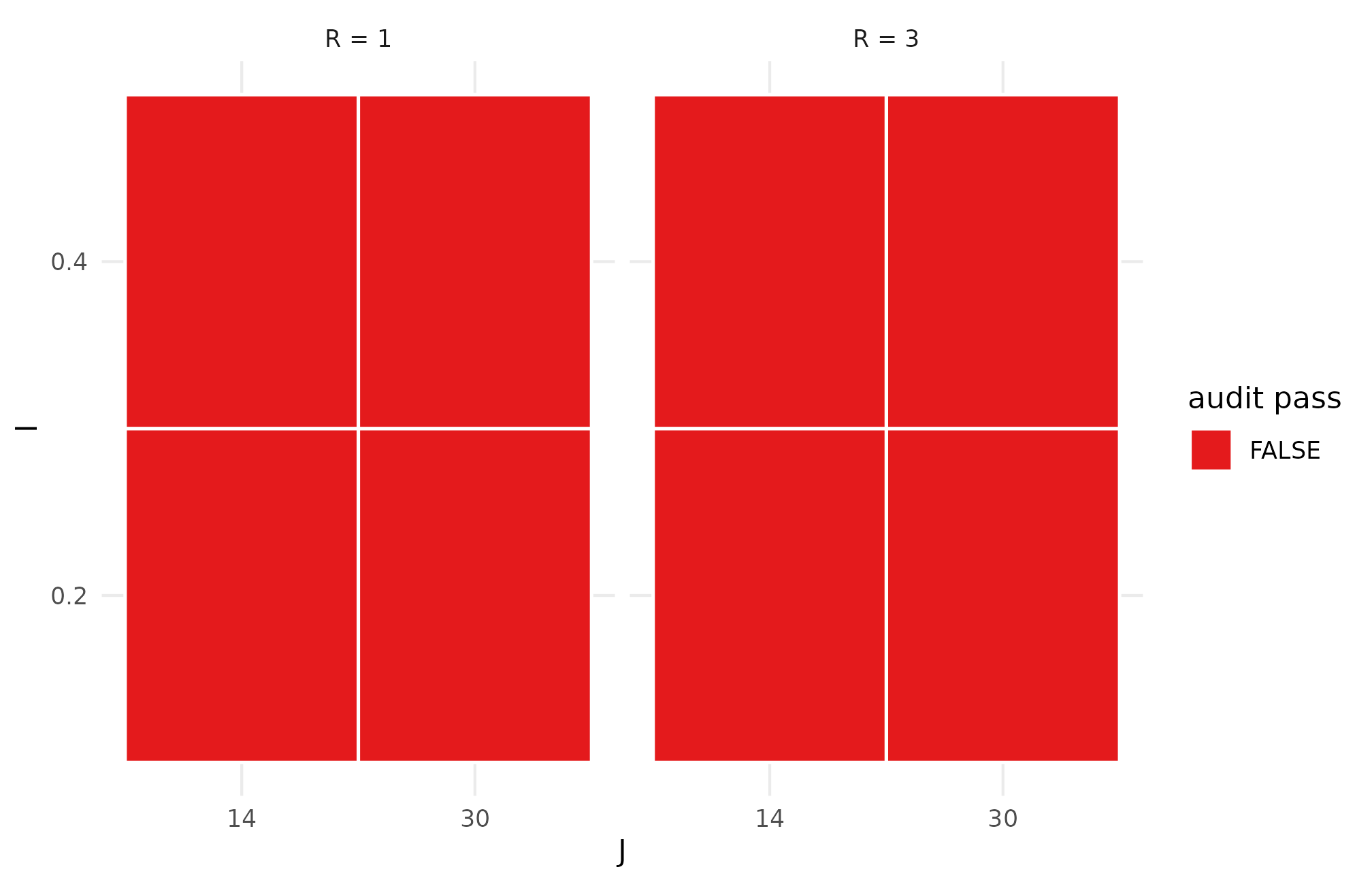
R5.7: scenario-audit pass/fail mosaic across the I x J grid faceted by R — green cells pass the diagnostic ladder at M = 1; the cells that fail concentrate where I is small and J is small, the regime where shrinkage cannot recover the between-site signal.
M = 1 is a smoke setting — for a defensible
pre-registration plan on M >= 50 and read off
med_I_hat, med_feasibility_efron, and the tail
quantiles. The longer narrative is in A6.
10. R5.8 — Canonical hash reproducibility
What it does: computes the canonical hash of the same design three times and confirms a single unique value, then permutes column order and confirms the hash is invariant.
When to reach for it: provenance and cache invalidation — a hash change on what should be the same artifact is a determinism signal worth chasing before publication.
library(multisiteDGP)
hashes <- vapply(
seq_len(3),
function(i) canonical_hash(sim_multisite(preset_education_modest(), seed = 12345L)),
character(1)
)
dat <- sim_multisite(preset_education_modest(), seed = 12345L)
reordered <- dat[, sort(names(dat))]
stopifnot(length(unique(hashes)) == 1L)
stopifnot(identical(canonical_hash(dat), canonical_hash(reordered)))
cat(
sprintf(
"COOKBOOK_RESULT R5.8 status=PASS unique_hashes=%d hash=%s\n",
length(unique(hashes)),
hashes[[1L]]
)
)Run it:
hashes_r58 <- vapply(
seq_len(3),
function(i) canonical_hash(
sim_multisite(preset_education_modest(), seed = 12345L)
),
character(1)
)
dat_r58 <- sim_multisite(preset_education_modest(), seed = 12345L)
reordered_r58 <- dat_r58[, sort(names(dat_r58))]
cat(sprintf(
"COOKBOOK_RESULT R5.8 status=PASS unique_hashes=%d hash=%s\n",
length(unique(hashes_r58)),
hashes_r58[[1L]]
))
#> COOKBOOK_RESULT R5.8 status=PASS unique_hashes=1 hash=7b960a5122df9c88The three reruns produce one unique hash:
hashes_r58
#> [1] "7b960a5122df9c88" "7b960a5122df9c88" "7b960a5122df9c88"And the column-permuted dataset hashes to the same value as the original, by design — the hash canonicalizes column order before serialization:
identical(canonical_hash(dat_r58), canonical_hash(reordered_r58))
#> [1] TRUETo make the determinism story visible, sweep across seeds and read off the per-seed hash:
hash_sweep_r58 <- data.frame(
seed = 12345L + 0:5,
hash = vapply(
12345L + 0:5,
function(s) canonical_hash(
sim_multisite(preset_education_modest(), seed = s)
),
character(1)
)
)
hash_sweep_r58
#> seed hash
#> 1 12345 7b960a5122df9c88
#> 2 12346 2058d5d7bb7d8fb8
#> 3 12347 92153f4d161ab874
#> 4 12348 0910f44251e84b16
#> 5 12349 94754d1e6036ad00
#> 6 12350 8aac98183a327912
hash_sweep_r58$short <- substr(hash_sweep_r58$hash, 1, 8)
ggplot(hash_sweep_r58, aes(factor(seed), 1, fill = short)) +
geom_tile(colour = "white", linewidth = 0.6) +
geom_text(aes(label = short), size = 3) +
scale_fill_grey(start = 0.6, end = 0.95) +
guides(fill = "none") +
labs(x = "seed", y = "") +
theme_minimal(base_size = 11) +
theme(axis.text.y = element_blank(),
axis.ticks.y = element_blank())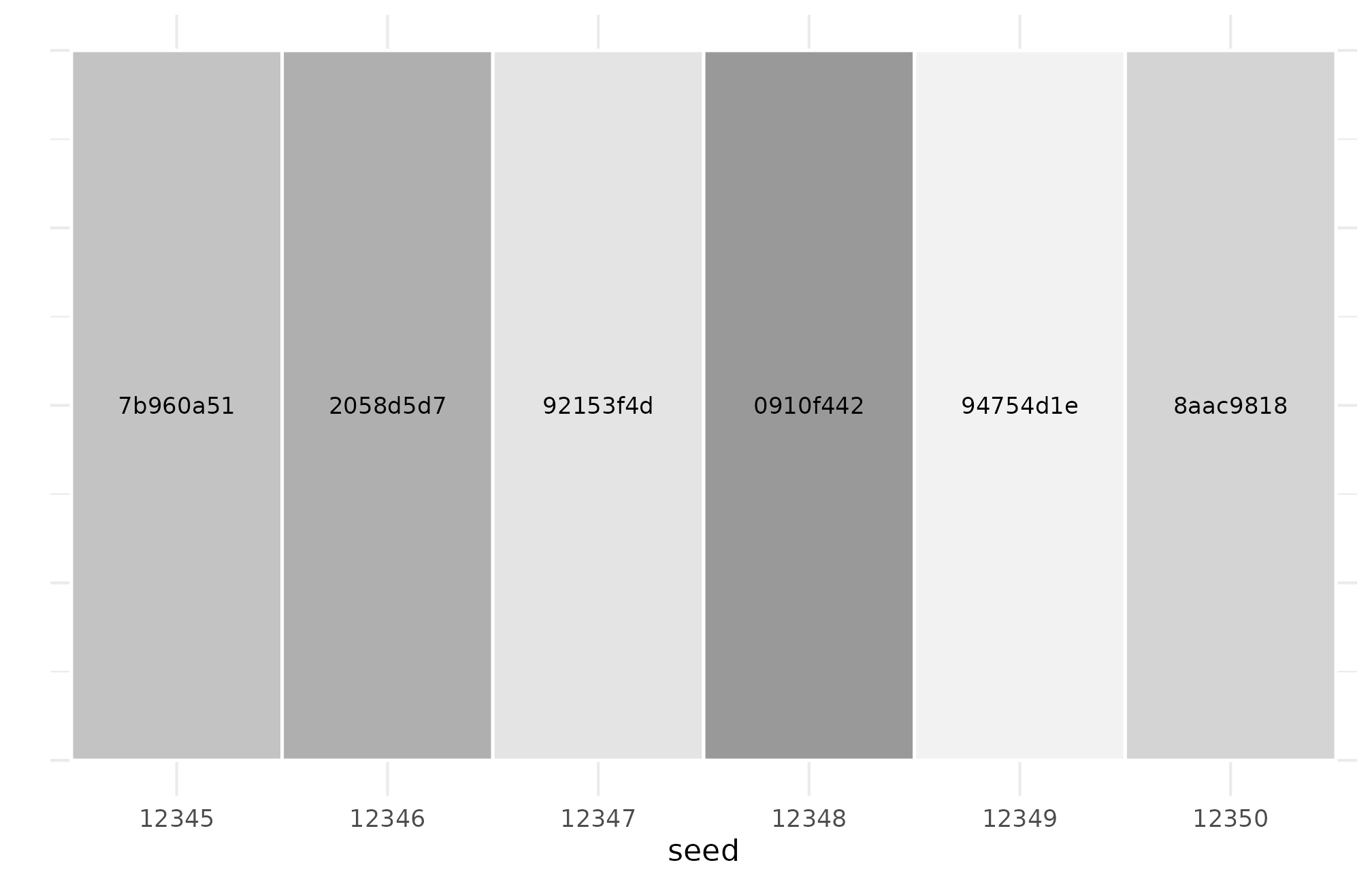
R5.8: per-seed hash uniqueness across six adjacent seeds — every seed produces a distinct hash, while same-seed reruns produce one stable hash, the two halves of canonical determinism.
The provenance idiom is one line:
provenance_string(dat_r58)
#> [1] "multisiteDGP 0.1.1 | paradigm=site_size | seed=12345 | canonical_hash=7b960a5122df9c88 | design_hash=db50cfd4e1659537 | hash_algo=xxhash64 | R=4.6.0 | hooks=none"That string is the artifact a reviewer can pin to a methods appendix (Lee et al., 2025).
11. R5.9 — Debugging feasibility diagnostics
What it does: simulates preset_education_modest() at
,
— small-trial conditions where the feasibility diagnostic does real work
— and reads informativeness() and
feasibility_index() across three adjacent seeds.
When to reach for it: a flagged feasibility warning. The first question is whether it is the design or the seed; this recipe is the shortest sanity check.
library(multisiteDGP)
design <- preset_education_modest(J = 14L, nj_mean = 60)
dat <- sim_multisite(design, seed = 12345L)
checks <- data.frame(
seed = 12345L + 0:2,
I = vapply(12345L + 0:2, function(s) informativeness(sim_multisite(design, seed = s)), numeric(1)),
feasibility = vapply(
12345L + 0:2,
function(s) feasibility_index(sim_multisite(design, seed = s), warn = FALSE),
numeric(1)
)
)
stopifnot(is.finite(informativeness(dat)))
stopifnot(is.finite(mean_shrinkage(dat)))
stopifnot(all(is.finite(checks$I)))
stopifnot(all(is.finite(checks$feasibility)))
cat(
sprintf(
"COOKBOOK_RESULT R5.9 status=PASS J=%d mean_I=%.3f mean_feasibility=%.3f\n",
design$J,
mean(checks$I),
mean(checks$feasibility)
)
)Run it:
design_r59 <- preset_education_modest(J = 14L, nj_mean = 60)
dat_r59 <- sim_multisite(design_r59, seed = 12345L)
checks_r59 <- data.frame(
seed = 12345L + 0:2,
I = vapply(
12345L + 0:2,
function(s) informativeness(sim_multisite(design_r59, seed = s)),
numeric(1)
),
feasibility = vapply(
12345L + 0:2,
function(s) feasibility_index(
sim_multisite(design_r59, seed = s), warn = FALSE
),
numeric(1)
)
)
cat(sprintf(
"COOKBOOK_RESULT R5.9 status=PASS J=%d mean_I=%.3f mean_feasibility=%.3f\n",
design_r59$J,
mean(checks_r59$I),
mean(checks_r59$feasibility)
))
#> COOKBOOK_RESULT R5.9 status=PASS J=14 mean_I=0.360 mean_feasibility=5.149The per-seed table is the actual debugging artifact:
checks_r59
#> seed I feasibility
#> 1 12345 0.3574726 5.096591
#> 2 12346 0.3566578 5.109289
#> 3 12347 0.3655686 5.241031The three seeds produce and feasibility values that move by under 5%, which is what you want — the feasibility warning is a property of the design, not the seed. If jumped by an order of magnitude across adjacent seeds, the design’s seed-to-seed sensitivity would be the diagnosis. Plot the variability:
ggplot(checks_r59, aes(factor(seed), feasibility)) +
geom_col(fill = "grey60", colour = "grey20") +
geom_text(aes(label = sprintf("%.2f", feasibility)),
vjust = -0.4, size = 3.5) +
labs(x = "seed", y = "feasibility index") +
theme_minimal(base_size = 11) +
expand_limits(y = max(checks_r59$feasibility) * 1.10)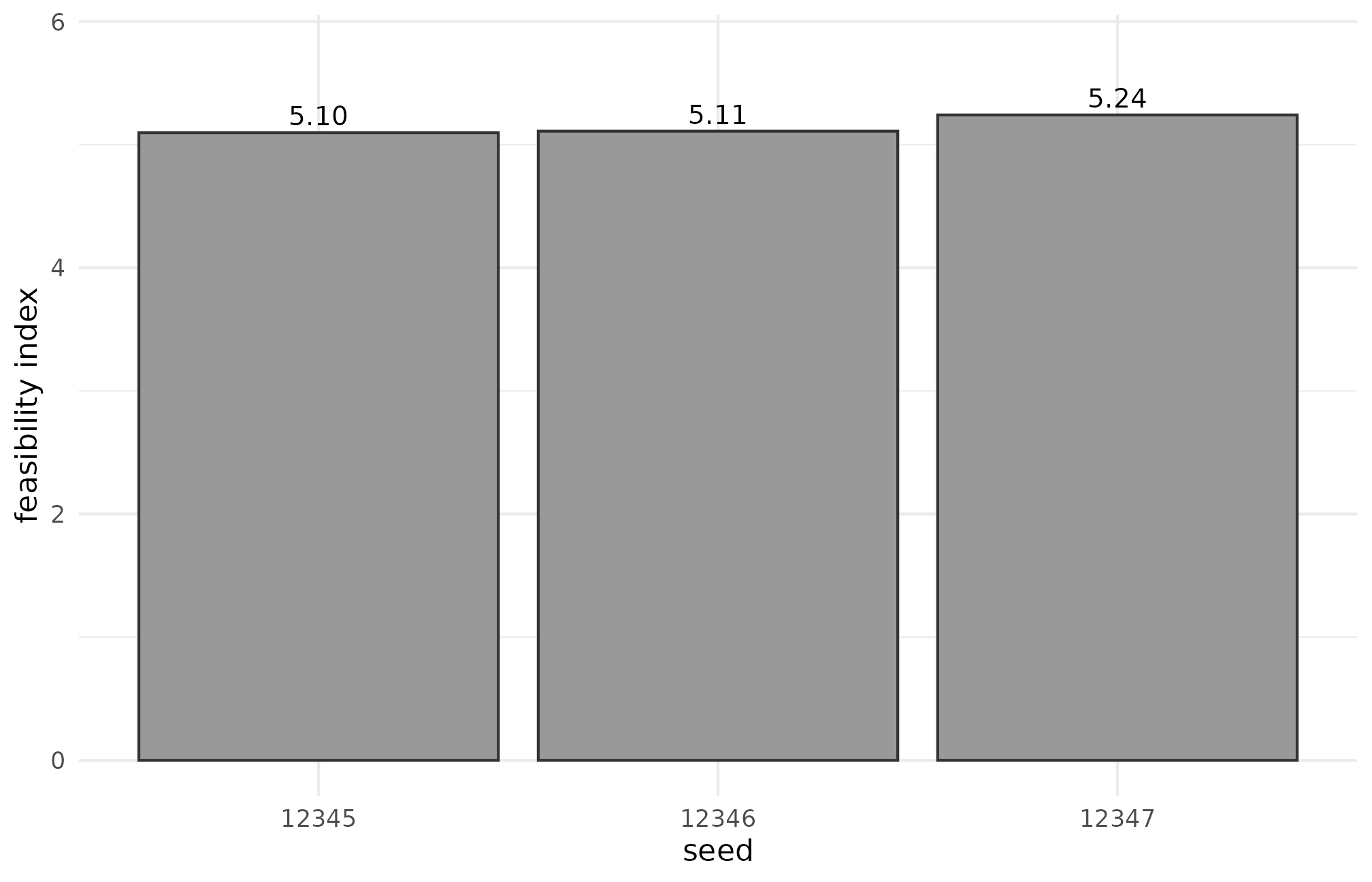
R5.9: feasibility index across three adjacent seeds at J = 14, n_j = 60 — bar heights are within 5% of each other, indicating the warning is a stable property of the design rather than a seed-specific artifact.
A small-trial regime (, ) is where the feasibility diagnostic earns its keep — diagnostics in practice are covered in A3.
12. The audit harness
The same harness that runs in CI is exposed for local use:
audit_script <- system.file(
"scripts", "cookbook_audit.R", package = "multisiteDGP"
)
env <- new.env(parent = globalenv())
sys.source(audit_script, envir = env)
env$run_cookbook_audit(execute = FALSE, verbose = FALSE)
#> id title file
#> 1 R5.1 Hello-world JEBS-style workflow R5-1-hello-world.R
#> 2 R5.2 Custom calibration with sigma_tau and R2 R5-2-custom-calibration.R
#> 3 R5.3 Compare two presets side by side R5-3-compare-presets.R
#> 4 R5.4 Bridge to downstream adapters R5-4-downstream-adapters.R
#> 5 R5.5 User g_fn callback R5-5-user-g-fn.R
#> 6 R5.6 Custom dependence_fn R5-6-custom-dependence-fn.R
#> 7 R5.7 Design grid and scenario audit R5-7-design-grid-audit.R
#> 8 R5.8 Canonical hash reproducibility R5-8-reproducibility-hash.R
#> 9 R5.9 Debugging feasibility diagnostics R5-9-debugging-feasibility.R
#> parsed executed marker_found error output_lines
#> 1 TRUE FALSE TRUE <NA> 0
#> 2 TRUE FALSE TRUE <NA> 0
#> 3 TRUE FALSE TRUE <NA> 0
#> 4 TRUE FALSE TRUE <NA> 0
#> 5 TRUE FALSE TRUE <NA> 0
#> 6 TRUE FALSE TRUE <NA> 0
#> 7 TRUE FALSE TRUE <NA> 0
#> 8 TRUE FALSE TRUE <NA> 0
#> 9 TRUE FALSE TRUE <NA> 0execute = FALSE parses the manifest and confirms every
recipe is discoverable; execute = TRUE runs each recipe
end-to-end and matches the printed marker against
expected_marker. A failed match is a smoke-test failure —
the audit harness is the canonical definition of “the recipes still
work”.
13. Where to next
The case studies pick up where the recipes stop, and the M-track vignettes document the four-layer mechanics behind every recipe.
- A6 · Case study — multisite trial — full pre-registration flow for a multisite trial: anchor design, scenario grid, audit, and methods-appendix template.
- A7 · Case study — meta-analysis — meta-analysis analogue: forest, funnel, dependence, and a heterogeneity-ratio narrative.
- M1 · The two-stage DGP — formal DGP behind every recipe; the canonical-column factorization.
- M3 · Margin and SE models — Layer 2 mechanics (site sizes and standard errors) that drive R5.4 / R5.7 / R5.9.
- M5 · Custom G distributions — full detail on the custom-callback contract used in R5.5 and R5.6.
- M7 · Reproducibility and provenance — hash + provenance machinery behind R5.8.
Acknowledgments
This research was supported by the Institute of Education Sciences, U.S. Department of Education, through Grant R305D240078 to the University of Alabama. The opinions expressed are those of the authors and do not represent views of the Institute or the U.S. Department of Education.
Session info
#> R version 4.6.0 (2026-04-24)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ggplot2_4.0.3 multisiteDGP_0.1.1
#>
#> loaded via a namespace (and not attached):
#> [1] gtable_0.3.6 xfun_0.57 bslib_0.10.0
#> [4] QuickJSR_1.9.2 ggrepel_0.9.8 inline_0.3.21
#> [7] lattice_0.22-9 mathjaxr_2.0-0 numDeriv_2016.8-1.1
#> [10] vctrs_0.7.3 tools_4.6.0 generics_0.1.4
#> [13] yulab.utils_0.2.4 parallel_4.6.0 stats4_4.6.0
#> [16] tibble_3.3.1 pkgconfig_2.0.3 Matrix_1.7-5
#> [19] checkmate_2.3.4 ggplotify_0.1.3 RColorBrewer_1.1-3
#> [22] S7_0.2.2 desc_1.4.3 RcppParallel_5.1.11-2
#> [25] lifecycle_1.0.5 compiler_4.6.0 farver_2.1.2
#> [28] textshaping_1.0.5 codetools_0.2-20 forestplot_3.2.0
#> [31] htmltools_0.5.9 sass_0.4.10 bayesplot_1.15.0
#> [34] yaml_2.3.12 tidyr_1.3.2 pillar_1.11.1
#> [37] pkgdown_2.2.0 crayon_1.5.3 jquerylib_0.1.4
#> [40] cachem_1.1.0 StanHeaders_2.32.10 metadat_1.6-0
#> [43] abind_1.4-8 nlme_3.1-169 rstan_2.32.7
#> [46] tidyselect_1.2.1 digest_0.6.39 purrr_1.2.2
#> [49] dplyr_1.2.1 splines_4.6.0 labeling_0.4.3
#> [52] fastmap_1.2.0 grid_4.6.0 cli_3.6.6
#> [55] metafor_5.0-1 magrittr_2.0.5 patchwork_1.3.2
#> [58] loo_2.9.0 pkgbuild_1.4.8 withr_3.0.2
#> [61] scales_1.4.0 backports_1.5.1 rappdirs_0.3.4
#> [64] rmarkdown_2.31 matrixStats_1.5.0 gridExtra_2.3
#> [67] ragg_1.5.2 evaluate_1.0.5 knitr_1.51
#> [70] mgcv_1.9-4 baggr_0.8 nleqslv_3.3.7
#> [73] gridGraphics_0.5-1 rstantools_2.6.0 rlang_1.2.0
#> [76] Rcpp_1.1.1-1.1 glue_1.8.1 jsonlite_2.0.0
#> [79] R6_2.6.1 systemfonts_1.3.2 fs_2.1.0