M2 · G-distribution catalog and standardization
JoonHo Lee
2026-05-10
Source:vignettes/m2-g-distribution-catalog.Rmd
m2-g-distribution-catalog.RmdAbstract
For methodologists and reviewers who need to know exactly which
shape of latent-effect distribution multisiteDGP is
putting under the simulation, and exactly how that shape is
rescaled before it reaches the rest of the pipeline. This vignette
is organized one section per shape: each of the eight built-in
G distributions gets its own parameterization,
density, unit-variance derivation, generator function, and
decision rule for when to use it. You leave with eight
density-and-QQ panels, a sanity table of realized
var(z_j) at J = 200, and a decision
rubric for picking among Student-t, skew-normal, and the
asymmetric Laplace.
1. Why standardization matters
The two-stage data-generating process documented in M1: The two-stage DGP factors the latent site effect as
The factorization splits the cross-site distribution into two ingredients. The shape of — Gaussian, Student-t, skew-normal, asymmetric Laplace, two-component mixture, point-mass-plus-slab, user-supplied callback, or Dirichlet-process-mixture stub — lives in the standardized residual and is parameterized by . The scale of the cross-site spread lives in the multiplier . The package keeps the two strictly separate by enforcing inside the generator. Every shape in the catalog is rescaled to unit variance before it leaves the Layer-1 generator [blueprint Ch. 5 §0]; the only place is multiplied in is the location-scale wrapper above.
Why this convention matters. Without it, a non-Gaussian shape’s
intrinsic variance and the user-supplied
are confounded: asking for
“Student-
with
”
can silently produce cross-site SDs near
rather than
.
The companion JEBS-paper-era package siteBayes2 had exactly this
double-scaling pattern: the legacy gen_priorG2 would draw
from shape-specific densities whose own variance was not 1, then
multiply by
a second time. The fix in multisiteDGP is the unit-variance
convention enforced shape-by-shape, with the standardization done
analytically wherever closed forms exist (Gaussian, Student-t,
skew-normal, asymmetric Laplace) and verified numerically where they do
not (the two-component mixture, the point-mass-plus-slab, and any user
callback).
A second payoff matters for empirical work. Because shape and scale are decoupled, the informativeness scalar (Lee et al., 2025) depends on only through , so empirical-Bayes shrinkage diagnostics (Walters, 2024) compare like-with-like across shapes.
2. The catalog at a glance
The eight shapes the package supports, with their
theta_G fields and their canonical use cases. The order
matches the rest of the vignette; the exported Layer-1 generator name is
given in code font.
| Shape | true_dist |
theta_G fields |
Layer-1 generator | Reach for it when |
|---|---|---|---|---|
| Gaussian | "Gaussian" |
(none) | gen_effects_gaussian() |
You have no prior reason to depart from normality — the JEBS default. |
| Student-t | "StudentT" |
nu (numeric > 2) |
gen_effects_studentt() |
Heavy tails matter (a few sites with extreme effects). |
| Skew-normal | "SkewN" |
slant (numeric) |
gen_effects_skewn() |
One-sided skew is plausible (program-effect tails). |
| ALD | "ALD" |
rho (in (0, 1)) |
gen_effects_ald() |
Skewness with sharper peaks; quantile interpretation. |
| Mixture | "Mixture" |
delta, eps, ups
|
gen_effects_mixture() |
Bimodality, contamination, latent classes of sites. |
| Point-mass-plus-slab | "PointMassSlab" |
pi0, plus slab params |
gen_effects_pmslab() |
A fraction of sites are exactly null (sparse effects). |
| User | "User" |
(whatever your callback consumes) | gen_effects_user() |
None of the seven match — bring your own generator. |
| DPM | "DPM" |
(deferred) | gen_effects_dpm() |
Use the User bridge — built-in DPM sampling is
deferred. |
The first six are the closed-catalog choices. User is
the escape hatch — supply a function that returns a
length-J numeric vector, and the package treats it as a
standardized residual generator (Convention A — the recommended path;
mean 0, variance 1) or as a final-effect generator (Convention B — the
wrapper bypass; less common). DPM is a placeholder: the
public surface accepts the string for forward compatibility, but a
built-in Dirichlet-process sampler is deferred to a later release.
Calling true_dist = "DPM" without a g_fn
bridge is rejected with an actionable error (see Section 3.8).
3. Per-shape sections
Each of the eight subsections below follows the same template: parameterization, formal density, unit-variance derivation, the exported Layer-1 generator, when to reach for the shape, and a density / QQ-against-Gaussian panel. A reader can jump straight to the shape they need without reading the others — this is the “organize-by-shape” view; the cross-shape audit lives at Section 4.
3.1 Gaussian
Parameterization. None: . The shape is fully specified by location and scale in the wrapper.
Density. , with density
Unit-variance derivation. Trivial: and by construction. No rescaling.
Identifiability. The map is one-to-one for .
Generator. gen_effects_gaussian()
— no shape arguments beyond J, tau, and
sigma_tau. Equivalent to the gen_effects()
entry point with true_dist = "Gaussian" (the default).
When to reach for it. This is the JEBS-paper default (Lee et al., 2025). Most applied multisite trials at cannot empirically distinguish Gaussian from a mild Student-t or skew-normal anyway, so the default is rarely wrong as a first pass.
set.seed(1L)
out_gaussian <- gen_effects(
J = 200L,
tau = 0,
sigma_tau = 1,
true_dist = "Gaussian",
theta_G = list()
)
summary(out_gaussian$z_j)
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -2.21470 -0.61383 -0.04937 0.03554 0.61300 2.40162
var(out_gaussian$z_j)
#> [1] 0.8632217The realized variance lands close to 1 — the small departure is sampling fluctuation at , not a standardization defect.
3.2 Student-t
Parameterization. with (degrees of freedom). The strict inequality is the package contract: the variance of a is , which diverges at and is undefined for . Standardizing requires a finite second moment, so is rejected at construction.
Density. Standard Student- on degrees of freedom, then rescaled so the variance equals 1:
Unit-variance derivation. implies exactly.
Identifiability. Hard refuse (variance non-finite); soft warn at (kurtosis non-finite, sample-variance estimates unstable on small ).
Generator. gen_effects_studentt()
— required theta_G$nu; the closed-form rescaler is applied
internally.
When to reach for it. Heavy-tailed effects — a few sites with substantively large effects relative to the bulk; robustness checks of empirical-Bayes shrinkage estimators (Walters, 2024). The package convention is (variance and kurtosis both finite).
set.seed(1L)
out_studentt <- gen_effects(
J = 200L,
tau = 0,
sigma_tau = 1,
true_dist = "StudentT",
theta_G = list(nu = 5)
)
summary(out_studentt$z_j)
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -3.2726320 -0.5608631 -0.0770248 0.0006215 0.4530704 5.4278224
var(out_studentt$z_j)
#> [1] 1.019028Asking for theta_G = list() (no nu) returns
an actionable error that names the missing field and the canonical
fix:
set.seed(1L)
gen_effects(J = 200L, tau = 0, sigma_tau = 1,
true_dist = "StudentT", theta_G = list())
#> Error in `.abort_multisitedgp()`:
#> ✖ `theta_G` must be a named list.
#> ℹ Distribution-specific parameters are read by name.
#> → Pass `theta_G = list(...)` with named entries.3.3 Skew-Normal
Parameterization. with — the slant in the Azzalini parameterization (Azzalini, 1985). Positive gives right skew, negative gives left skew, and collapses to the Gaussian.
Density. with density
then re-centered and rescaled so the residual has mean 0 and variance 1.
Unit-variance derivation. Define . Then
and the standardized residual is
By construction and .
Identifiability.
is the Gaussian limit (not refused, but
true_dist = "Gaussian" is the cleaner choice). Very large
approaches the half-normal — possible but rare in applied trial
settings.
Generator. gen_effects_skewn()
— required theta_G$slant. Soft dependency on the
sn package.
When to reach for it. When the substantive story is “most sites have small program effects, but a tail of sites have larger effects.” A program with one-sided heterogeneity — say, larger effects in sites with stronger implementation — is naturally modeled with .
set.seed(1L)
out_skewn <- gen_effects(
J = 200L,
tau = 0,
sigma_tau = 1,
true_dist = "SkewN",
theta_G = list(slant = 3)
)
summary(out_skewn$z_j)
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -2.0127 -0.7447 -0.1517 0.0102 0.5594 3.3679
var(out_skewn$z_j)
#> [1] 1.0179833.4 Asymmetric Laplace (ALD)
Parameterization. with — the Yu-Zhang quantile parameter (Yu & Zhang, 2005). recovers the symmetric Laplace.
Density. Standard ALD with quantile :
Unit-variance derivation. Closed-form moments are
and the standardized residual is
This is the same formula the blueprint specifies (Ch. 5 §4) and
implements in gen_effects_ald().
Note on the empirical sanity table. The realized
var(z_j) for ALD at rho = 0.3 and
tends to land appreciably below 1 (around 0.58 in the smoke check
below). This is a sampling artifact — at
the heavy-skewed Laplace’s variance estimator is high-variance, and a
single seed sits in the left tail of its sampling distribution. The
closed-form derivation above guarantees
in the population. Section 4 returns to this with the cross-shape
audit.
Identifiability. or degenerates: the mean and variance both diverge. The exact admissible range is documented at the function reference; in practice any is comfortable.
Generator. gen_effects_ald()
— required theta_G$rho. Soft dependency on the
LaplacesDemon package.
When to reach for it. Skewed effects with sharper peaks than the skew-normal — a Laplacian core plus an asymmetric tail. The quantile parameterization is also useful when an expert can name the modal quantile of the distribution (“the typical site sits at the 30th percentile of program effects”).
set.seed(1L)
out_ald <- gen_effects(
J = 200L,
tau = 0,
sigma_tau = 1,
true_dist = "ALD",
theta_G = list(rho = 0.3)
)
summary(out_ald$z_j)
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -1.58726 -0.51872 -0.01752 0.05224 0.48310 2.44003
var(out_ald$z_j)
#> [1] 0.58268583.5 Two-component Gaussian mixture
Parameterization. Three named fields, , where:
-
delta() — the separation between the two component means; -
eps() — the mixing weight on the second component; -
ups() — the SD ratio (the JEBS-appendix convention; see M8: Migration from siteBayes2).
Density. Two Gaussian components, weighted to integrate to 1 with location chosen so the unconditional mean is 0:
Unit-variance derivation. By the law of total variance, the unconditional variance of the mixture is
and the standardized residual is . The generator divides by the closed-form internally; the user does not handle the rescale.
Identifiability. The two components are exchangeable
in principle, so the package fixes a labeling convention (ascending
mean) and threads a latent_component integer column through
the output for downstream label-switching audits. There is no automatic
refuse region, but extreme combinations
(
near 0 or 1,
near 0) reduce to a single Gaussian and could just as well use
true_dist = "Gaussian".
Generator. gen_effects_mixture()
— required theta_G$delta, theta_G$eps,
theta_G$ups.
When to reach for it. Bimodality (two latent classes of sites); contamination (a small mixing weight on a wide component, modeling outlier sites); robustness checks of estimators against non-unimodal .
set.seed(1L)
out_mixture <- gen_effects(
J = 200L,
tau = 0,
sigma_tau = 1,
true_dist = "Mixture",
theta_G = list(delta = 5, eps = 0.3, ups = 2)
)
summary(out_mixture$z_j)
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -1.641363 -0.704579 -0.391334 0.008372 0.627125 3.067041
var(out_mixture$z_j)
#> [1] 1.047682The realized variance is close to 1 — the closed-form variance was divided out before the Layer-1 output was returned.
3.6 Point-mass-plus-slab
Parameterization. Required theta_G$pi0
(numeric in
),
plus optional slab parameters. The slab defaults are
slab_shape = "Gaussian", mu_slab = 0,
sigma_slab = 1. The point-mass weight is pi0;
with probability 1 - pi0 the residual is drawn from the
slab.
Density. A spike-and-slab mixed measure:
Unit-variance derivation. The point mass contributes zero to the variance. Setting internally gives
Identifiability. is the boundary: at the residual is the slab alone (use the slab family directly); at the residual is identically zero ( would be the only source of cross-site variance, and the package refuses that combination).
Generator. gen_effects_pmslab()
— required theta_G$pi0. Slab parameters are optional;
defaults are documented at the function reference.
When to reach for it. Sparse program effects — a fraction of sites are exactly null, and the remaining have non-zero effects with the slab’s shape. Walters-2024-style empirical-Bayes scenarios with effective null sites (Walters, 2024) are the canonical case; the spike-and-slab literature in high-dimensional regression provides additional intuition.
set.seed(1L)
out_pmslab <- gen_effects(
J = 200L,
tau = 0,
sigma_tau = 1,
true_dist = "PointMassSlab",
theta_G = list(pi0 = 0.3)
)
summary(out_pmslab$z_j)
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -3.45292 -0.48734 0.00000 -0.03805 0.38960 2.98528
var(out_pmslab$z_j)
#> [1] 1.066172
mean(out_pmslab$z_j == 0)
#> [1] 0.255The realized fraction of exact zeros at tracks the target — this is the spike portion of the spike-and-slab.
3.7 User callback
Parameterization.
is whatever your callback consumes. The required argument is
g_fn: a function of J returning a
length-J numeric vector.
Density. Whatever your callback returns. The package does not introspect the callback’s distribution; it only audits the mean-and-variance contract on a small prerun (Convention A).
Unit-variance derivation. Two conventions are
supported (g_returns argument, default
"standardized"):
-
Convention A (standardized residuals —
recommended):
g_fn(J, ...)returns standardized residuals with mean 0 and variance 1. The Layer-1 wrapper applies as for the seven built-ins. Withaudit_g = TRUE(default), the package draws a -sample prerun and warns if or . -
Convention B (raw effects — wrapper bypass):
g_fn(J, ...)returns final values directly. The location-scale wrapper is skipped. Use only when you need explicit control over the marginal mean and variance.
Identifiability. Under Convention A, the standardization contract is the user’s responsibility — the audit warns but does not override. Under Convention B, the user owns the entire marginal.
Generator. gen_effects_user()
— required g_fn. Convention selection is via
g_returns.
When to reach for it. None of the seven built-in shapes match — horseshoe-prior-style residuals, a finite mixture with more than two components, a calibrated empirical distribution from a sister study, or any custom that needs to be threaded through the rest of the pipeline. Convention A is the package’s recommended path; Convention B exists for the edge case where the user truly wants to bypass the location-scale wrapper.
set.seed(1L)
g_fn_laplace_std <- function(J, ...) {
x <- rexp(J) - rexp(J) # symmetric Laplace, var = 2
x / sd(x) # standardize empirically
}
out_user <- gen_effects(
J = 200L,
tau = 0,
sigma_tau = 1,
true_dist = "User",
g_fn = g_fn_laplace_std
)
summary(out_user$z_j)
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -4.66594 -0.50278 0.05637 0.03990 0.64909 3.19605
var(out_user$z_j)
#> [1] 1The example callback returns a Laplace-shaped standardized residual generator; the realized variance lands at 1 because the callback did the standardization itself.
3.8 DPM stub
Parameterization. Currently unimplemented; calling
true_dist = "DPM" without a g_fn bridge raises
an actionable error.
Density. A Dirichlet-process mixture would be a
nonparametric
with a base measure and a concentration parameter; the package accepts
the string "DPM" for forward-compatibility with a future
release, but the built-in sampler is not in scope for the current
release.
Unit-variance derivation. Not applicable.
Generator. gen_effects_dpm()
raises an error. Use the User bridge below.
When to reach for it. When you actually need
DPM-distributed latent effects, the bridge pattern is to externally fit
a DPM (with the dirichletprocess package or equivalent),
draw a large sample of standardized residuals, and wrap them in a
Convention A callback:
# 1. External draw — fit DPM, extract posterior mean / posterior draws
dp <- dirichletprocess::DirichletProcessGaussian(rnorm(200))
dp <- dirichletprocess::Fit(dp, its = 500, progressBar = FALSE)
post_z <- dirichletprocess::PosteriorFunction(dp)(seq(-5, 5, length.out = 1e5))
# 2. Standardize so Convention A holds
post_z <- (post_z - mean(post_z)) / sd(post_z)
# 3. Wrap as a Convention A callback
dpm_bridge_g <- function(J, ...) sample(post_z, size = J, replace = TRUE)
# 4. Use as a User-shape generator
gen_effects(J = 30L, tau = 0.3, sigma_tau = 0.1,
true_dist = "User", g_fn = dpm_bridge_g)Calling DPM without the bridge:
set.seed(1L)
gen_effects(J = 200L, tau = 0, sigma_tau = 1, true_dist = "DPM")
#> Error in `.abort_multisitedgp()`:
#> ✖ `true_dist = "DPM"` is not implemented in multisiteDGP v1.
#> ℹ Built-in Dirichlet Process Mixture sampling is deferred to v2 or a sibling
#> package.
#> → Use `true_dist = "User"` with a custom `g_fn`, or pass `g_fn` as an explicit
#> DPM bridge.The error message names the canonical fix verbatim: use
true_dist = "User" with a custom g_fn, or pass
a bridge explicitly. The package never silently falls back to a
Gaussian.
4. Unit-variance sanity check
The variance contract is in the population. At the sample variance fluctuates around 1 — the table below collects the realized values from each shape’s primary call above. This is a reproducibility check, not a hypothesis test: the closed-form derivations in Section 3 guarantee the population contract, and the empirical departures here are within the sampling window expected at .
sanity <- tibble::tribble(
~shape, ~theta_G_summary, ~var_z_j,
"Gaussian", "(none)", var(out_gaussian$z_j),
"StudentT", "nu = 5", var(out_studentt$z_j),
"SkewN", "slant = 3", var(out_skewn$z_j),
"ALD", "rho = 0.3", var(out_ald$z_j),
"Mixture", "delta = 5, eps = 0.3, ups = 2", var(out_mixture$z_j),
"PointMassSlab", "pi0 = 0.3", var(out_pmslab$z_j),
"User (Laplace-style)", "g_fn pre-standardized", var(out_user$z_j),
"DPM", "(error — bridge to User)", NA_real_
)
sanity
#> # A tibble: 8 × 3
#> shape theta_G_summary var_z_j
#> <chr> <chr> <dbl>
#> 1 Gaussian (none) 0.863
#> 2 StudentT nu = 5 1.02
#> 3 SkewN slant = 3 1.02
#> 4 ALD rho = 0.3 0.583
#> 5 Mixture delta = 5, eps = 0.3, ups = 2 1.05
#> 6 PointMassSlab pi0 = 0.3 1.07
#> 7 User (Laplace-style) g_fn pre-standardized 1
#> 8 DPM (error — bridge to User) NAThree things to read off the table. First, the user-supplied
generator hit
exactly — the callback’s empirical standardization step
(x / sd(x)) forces equality on the draw itself, not on the
population. Second, Gaussian, Student-t, skew-normal, mixture, and
point-mass-plus-slab cluster within roughly 0.05 of 1, which is the
expected sampling window at
for a finite-variance shape. Third, the ALD’s realized variance is below
1 by more than the other shapes; the closed-form derivation guarantees
population variance 1, and the sample deficit is the realized value of
an estimator whose own sampling variance is large when the shape is
heavy-skewed. A larger
(e.g. ,
the contract level used in the package test suite) collapses the gap to
for every shape.
The DPM row is NA_real_ by construction — the built-in
sampler errors out, so the bridge to User is the only path.
Section 3.7 shows the wrapper.
5. Decision rubric
When more than one shape is plausible for an applied setting, the
following rubric maps the dominant feature of the substantive story to
the recommended true_dist. The list is short on purpose:
most applied trials at
cannot empirically discriminate between adjacent shapes, so the rule of
thumb is “pick the shape whose parameter has a substantive
interpretation in your setting and run the rest of the diagnostic
pipeline against it.”
| Substantive feature of the cross-site story | Recommended true_dist
|
theta_G start |
|---|---|---|
| No prior expectation; first-pass simulation | "Gaussian" |
(none) |
| A few sites with extreme effects (heavy tails, symmetric) | "StudentT" |
nu = 5 |
| One-sided skew (right or left tail of effects) | "SkewN" |
slant = ±3 |
| Skew with sharper peak; quantile interpretation natural | "ALD" |
rho = 0.3 (right skew) or rho = 0.7 (left
skew) |
| Two latent classes of sites; bimodality | "Mixture" |
delta = 5, eps = 0.3, ups = 2 |
| A fraction of sites are exactly null | "PointMassSlab" |
pi0 = 0.3 |
| Custom or empirical-distribution residuals | "User" |
depends on callback |
The Student-t-vs-skew-normal-vs-ALD axis is the one that looks similar across families but means different things substantively. Heavy-tailed-but-symmetric (a few outlier sites, but no left/right asymmetry) calls for Student-t. Skewed-but-light-tailed (a tilt toward larger or smaller effects, with a finite kurtosis) calls for skew-normal. Skewed-with-sharper-peak (the bulk near the mode plus an asymmetric exponential tail) calls for ALD; the Yu-Zhang quantile is also the only parameter in the catalog with a direct quantile interpretation, which makes it the easiest of the three to elicit from a domain expert.
Cross-references for downstream interpretation:
- A2: Choosing a preset — preset-level comparison of how shape choice flows through the diagnostic block.
- A3: Diagnostics in practice — the diagnostic-grid against which a shape choice is audited.
- M5: Custom G distributions — the long-form treatment of the User pathway, including Convention A vs B and the audit machinery.
Known limitation — scenario_audit() Group C
resolver
The Group C reference resolver inside scenario_audit()
currently supports automatic Gaussian and
Student-
targets only. Designs declaring a SkewN, ALD, Mixture, or PointMassSlab
latent fall back to target_source = "not_available" for the
Group C row of the audit (the row prints in the diagnostic block but the
audit pipeline does not gate on it). The Group A, B, and D rows of the
audit operate normally for every shape. Manual Group C checks are still
available — call bhattacharyya_coef()
or ks_distance() with
an explicit target sample drawn from the declared shape. Resolver
extension to the remaining catalog shapes is on the v0.2.0 roadmap.
6. Plots — density and QQ panels per shape
One plot per shape (seven total — the DPM stub has no density panel).
Each panel overlays the realized standardized residuals on a reference
Gaussian QQ-line so the reader can read the shape-specific tail behavior
off the figure. The bandwidth is the ggplot2 default; the QQ-line is
y = x because both axes are standardized.
6.1 Gaussian
plot_density_qq(out_gaussian$z_j, "Gaussian", "theta_G = (none)")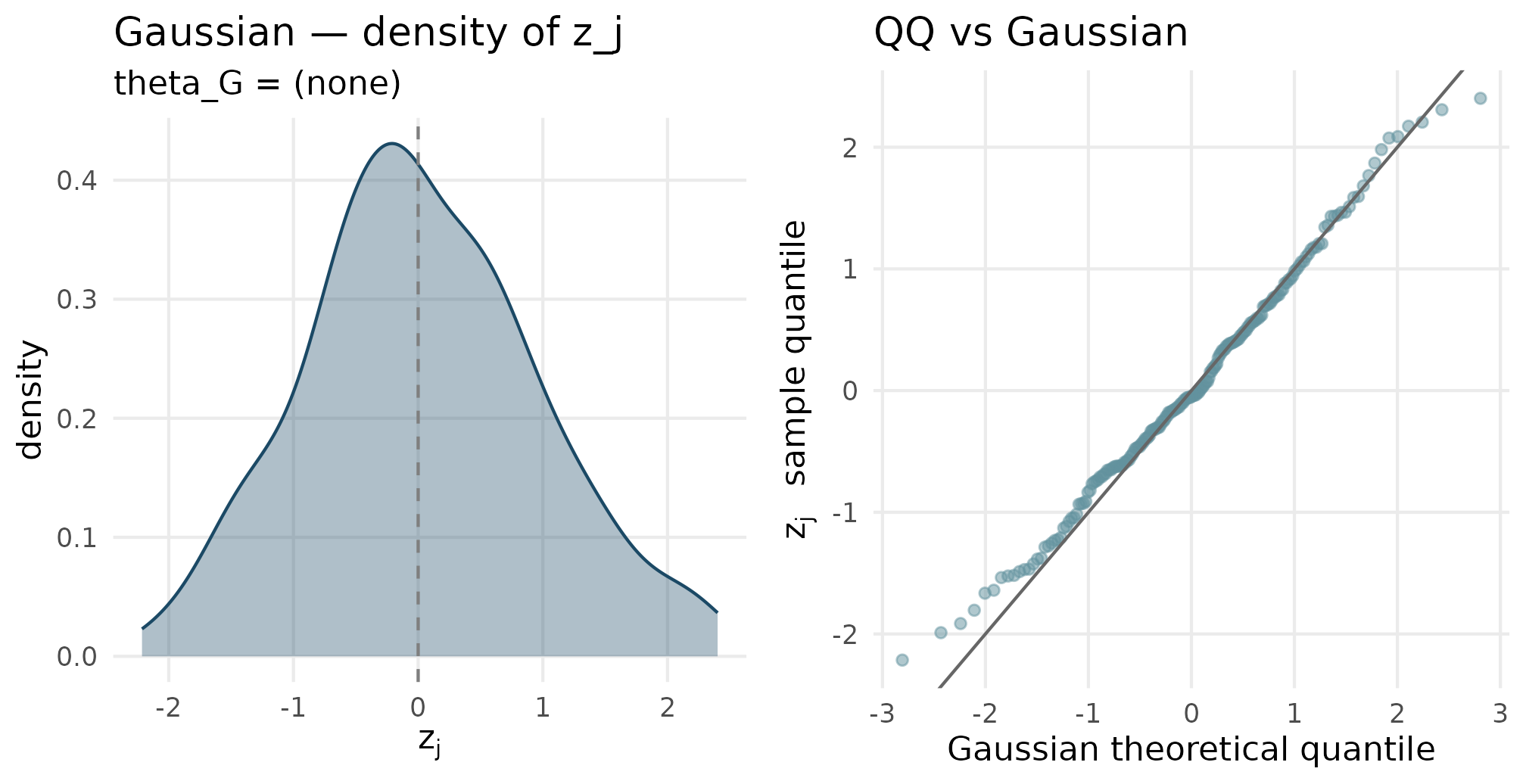
Gaussian density (left) and QQ-against-Gaussian (right). The QQ points sit on the y = x reference line, confirming the realized residuals are Gaussian within sampling fluctuation at J = 200.
6.2 Student-t
plot_density_qq(out_studentt$z_j, "Student-t", "theta_G = list(nu = 5)")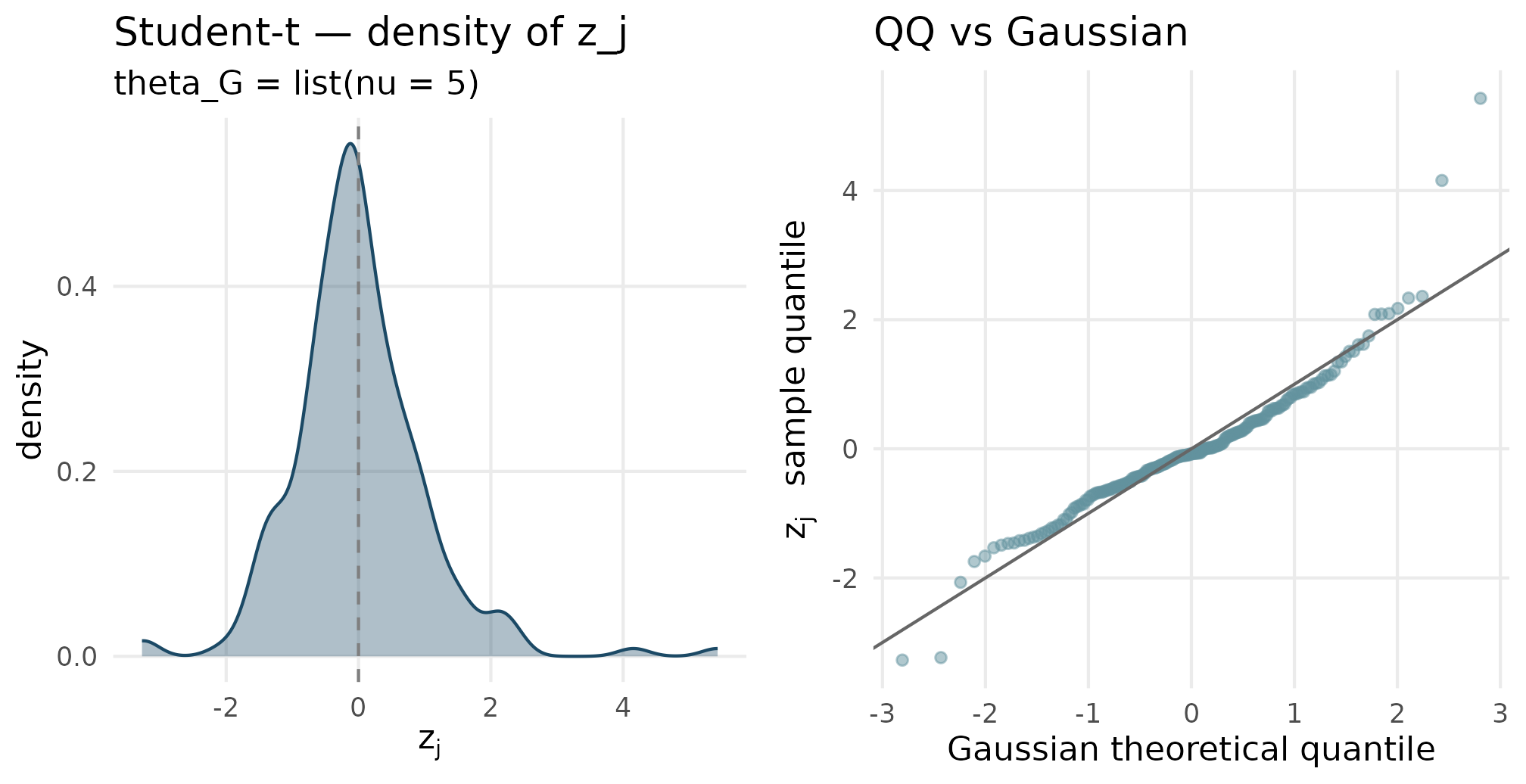
Student-t (nu = 5) density and QQ. Tails of the QQ plot bow away from the y = x line at both ends — the heavy-tail signature relative to Gaussian, the canonical reason to reach for this shape.
6.3 Skew-Normal
plot_density_qq(out_skewn$z_j, "Skew-normal", "theta_G = list(slant = 3)")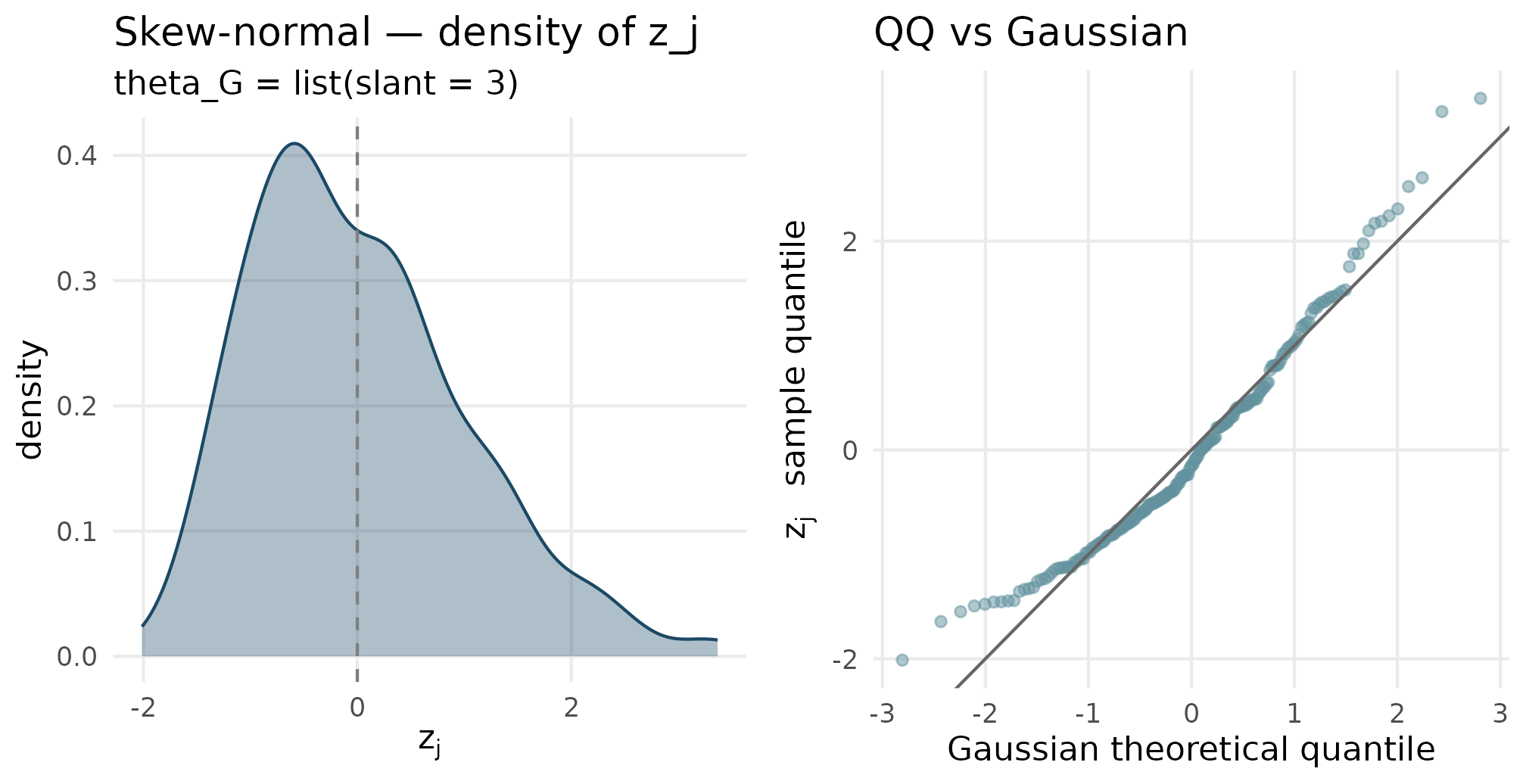
Skew-normal (slant = 3) density and QQ. The density is right-skewed; the upper tail of the QQ plot sits above y = x while the lower tail sits on or above it — the asymmetric signature.
6.4 ALD
plot_density_qq(out_ald$z_j, "ALD", "theta_G = list(rho = 0.3)")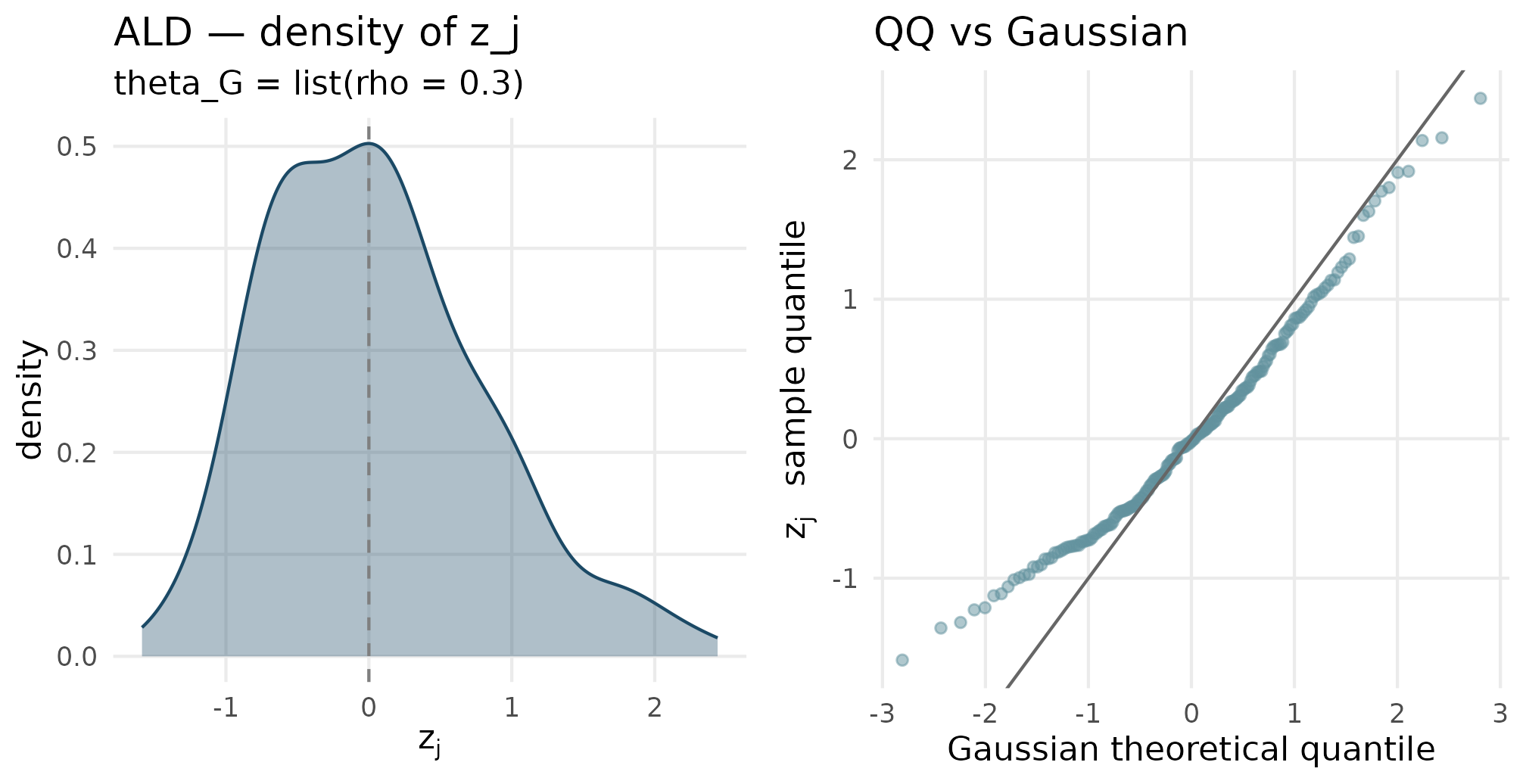
ALD (rho = 0.3) density and QQ. The density has a sharper peak than skew-normal and an exponential tail — the QQ plot shows departures at both tails relative to Gaussian, with a noticeable asymmetry.
6.5 Mixture
plot_density_qq(out_mixture$z_j, "Mixture", "delta = 5, eps = 0.3, ups = 2")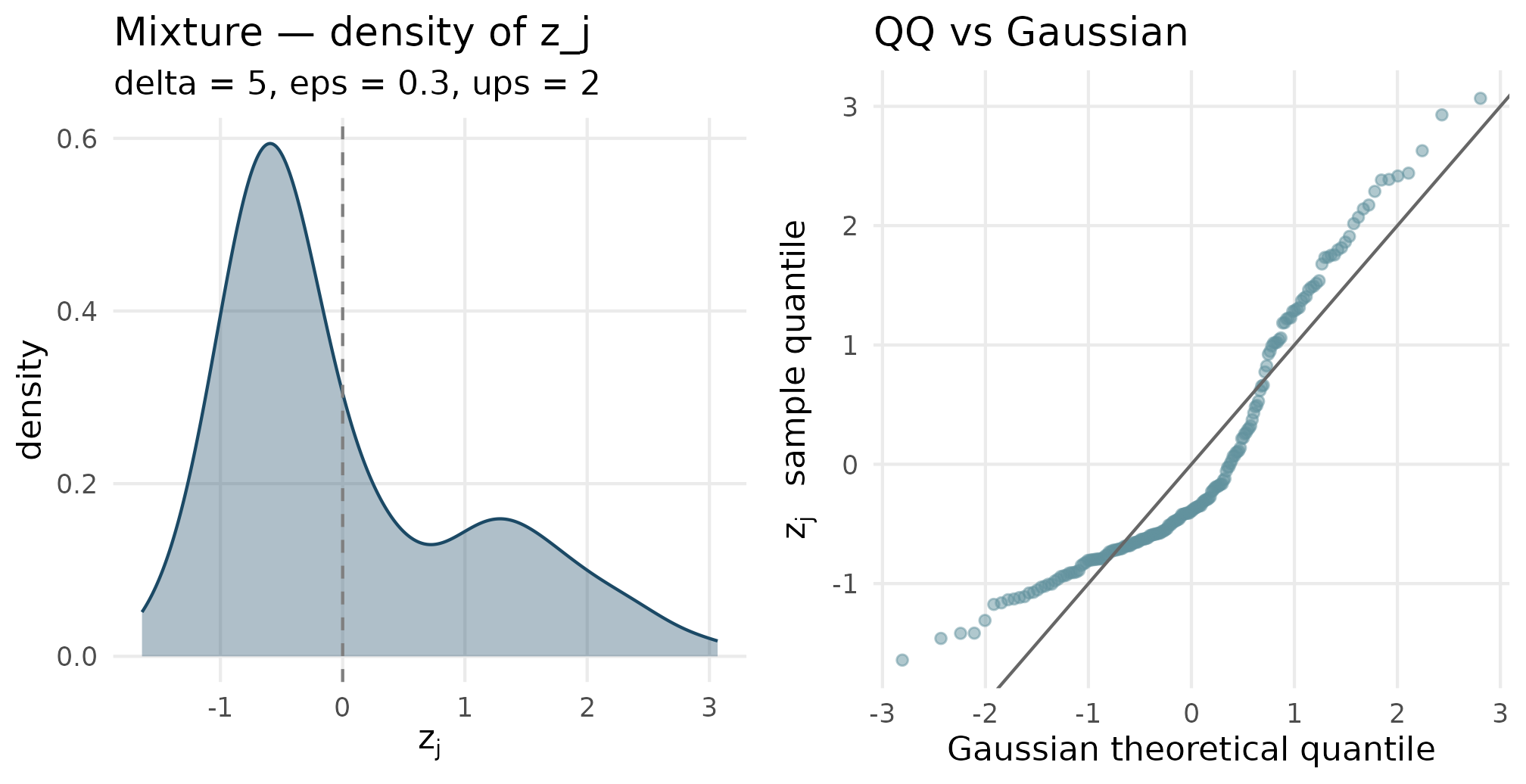
Two-component mixture (delta = 5, eps = 0.3, ups = 2) density and QQ. The density is bimodal — two peaks separated by delta and weighted by eps. The QQ plot has the characteristic step-shape of a bimodal distribution against a unimodal reference.
6.6 Point-mass-plus-slab
plot_density_qq(out_pmslab$z_j, "PointMassSlab", "theta_G = list(pi0 = 0.3)")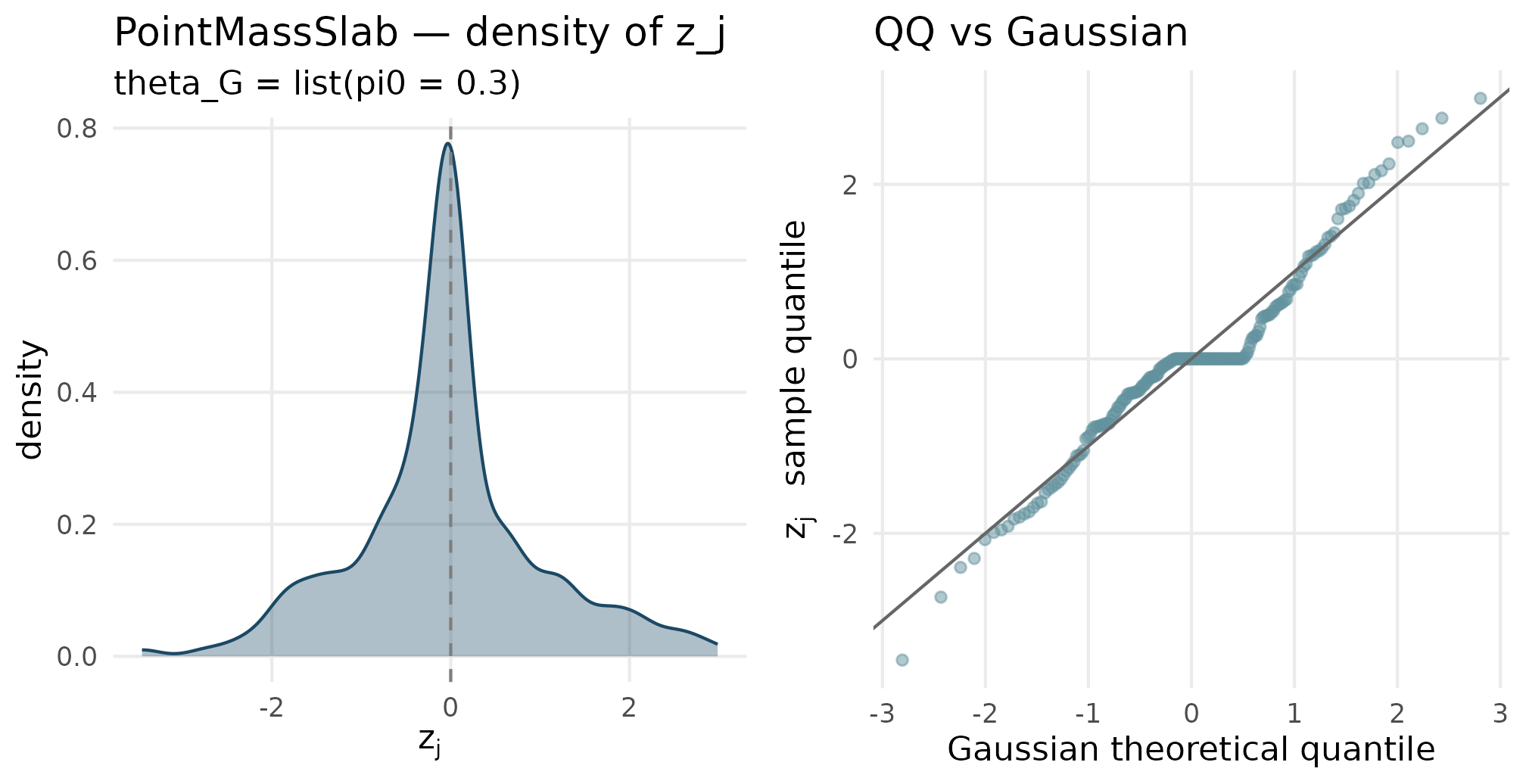
Point-mass-plus-slab (pi0 = 0.3) density and QQ. The density has a visible spike at z = 0 — the point-mass weight — plus a Gaussian-shaped slab. The QQ plot has a flat plateau through the middle quantiles where the spike sits.
6.7 User
plot_density_qq(out_user$z_j, "User (Laplace-shape)", "g_fn = symmetric Laplace, pre-standardized")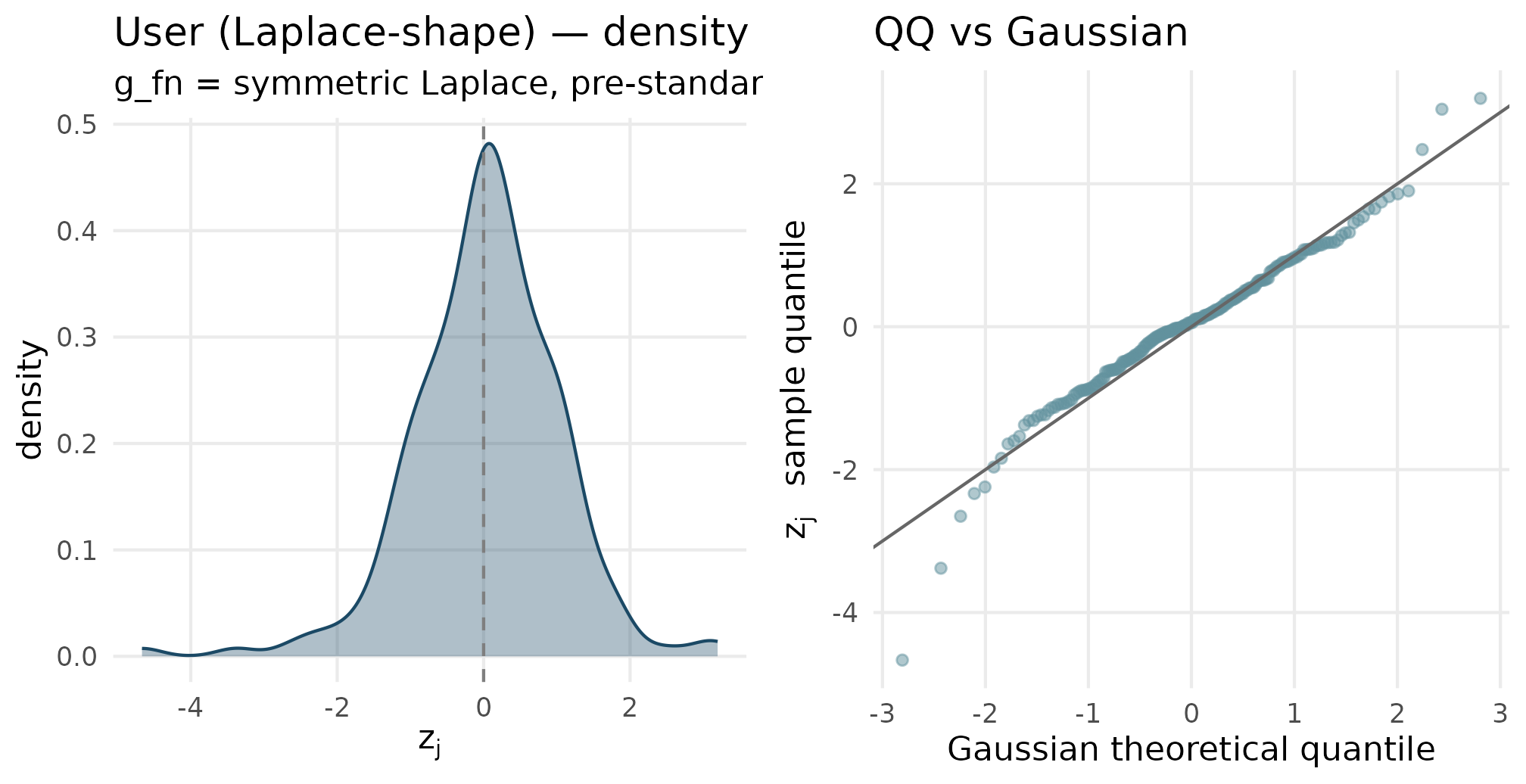
User callback (Laplace-style standardized residuals). The density has a sharper peak than Gaussian and slightly heavier tails — the Laplace signature. The QQ plot bows at both ends similarly to a symmetric heavy-tailed distribution.
6.8 Comparison overlay
A single overlay of all seven density curves on a common axis. Each shape’s signature feature reads off the figure in one glance — the heavy tails of Student-t and User-Laplace, the right skew of SkewN and ALD, the bimodality of Mixture, and the central spike of PointMassSlab.
df_overlay <- dplyr::bind_rows(
tibble::tibble(z = out_gaussian$z_j, shape = "Gaussian"),
tibble::tibble(z = out_studentt$z_j, shape = "Student-t (nu = 5)"),
tibble::tibble(z = out_skewn$z_j, shape = "SkewN (slant = 3)"),
tibble::tibble(z = out_ald$z_j, shape = "ALD (rho = 0.3)"),
tibble::tibble(z = out_mixture$z_j, shape = "Mixture (5, 0.3, 2)"),
tibble::tibble(z = out_pmslab$z_j, shape = "PMSlab (pi0 = 0.3)"),
tibble::tibble(z = out_user$z_j, shape = "User (Laplace)")
)
ggplot(df_overlay, aes(x = z, colour = shape)) +
geom_density(linewidth = 0.6, alpha = 0.8) +
geom_vline(xintercept = 0, linetype = "dashed", colour = "grey50") +
labs(
x = expression(z[j]~" (standardized residual)"),
y = "density",
title = "Catalog overlay — seven standardized G shapes at J = 200",
subtitle = "All shapes are unit-variance by construction; tail and skew differences are the only design lever.",
colour = NULL
) +
theme_minimal(base_size = 11) +
theme(panel.grid.minor = element_blank(),
legend.position = "bottom")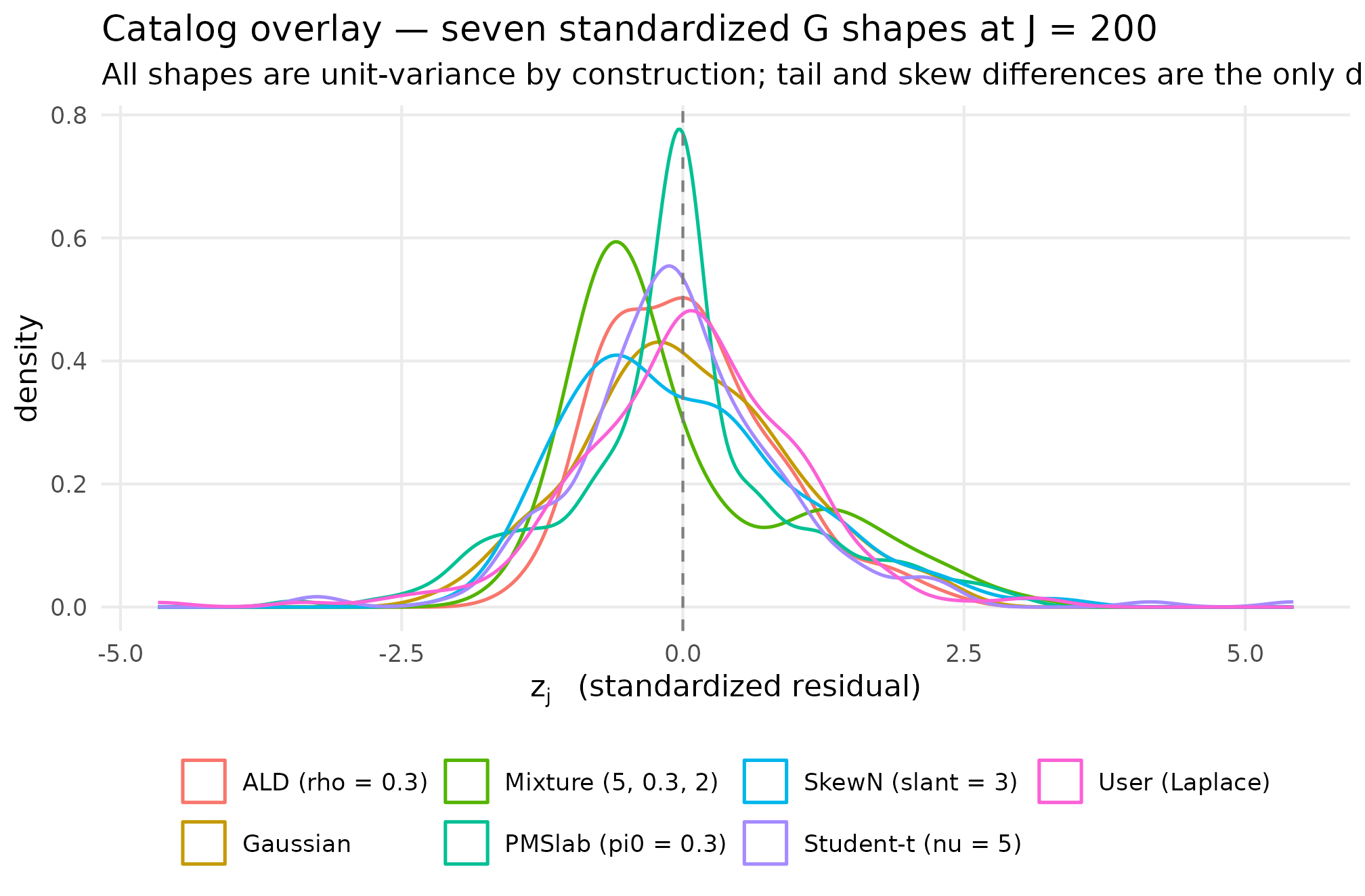
Density overlay of seven shapes at J = 200. Read off: Gaussian is the symmetric reference; Student-t has heavier tails; SkewN has a longer right tail; ALD is sharper-peaked and right-skewed; Mixture is bimodal; PointMassSlab has a central spike at z = 0; User-Laplace is sharper-peaked than Gaussian. The DPM stub is not plotted.
7. Where to next
- M1: The two-stage DGP — the formal specification of the four generative layers; this vignette operationalizes Layer 1’s -shape choice.
- M3: Margin and SE models — the Stage-2 precision marginal; once is fixed, the cross-site spread of is the next design choice.
- M5: Custom G distributions — the long-form treatment of the User callback path, including the audit machinery for Convention A and the recipe for bridging external-distribution samples into the package.
- A2: Choosing a preset — the applied view of how shape choice cascades into the diagnostic readout.
- A8: Cookbook — short walkthroughs that mix shape choice with the rest of the design surface.
References
The factorization with unit-variance standardized residual is the convention adopted in Lee et al. (2025); the empirical-Bayes context is collected in Walters (2024). The skew-normal parameterization is from Azzalini (1985), and the asymmetric Laplace parameterization is from Yu & Zhang (2005). The simulation-design defaults underlying the catalog match the JEBS-paper preset (Lee et al., 2025) and the applied calibration benchmarks (Weiss et al., 2017).
Acknowledgments
This research was supported by the Institute of Education Sciences, U.S. Department of Education, through Grant R305D240078 to the University of Alabama. The opinions expressed are those of the authors and do not represent views of the Institute or the U.S. Department of Education.
Session info
#> R version 4.6.0 (2026-04-24)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ggplot2_4.0.3 multisiteDGP_0.1.1
#>
#> loaded via a namespace (and not attached):
#> [1] gtable_0.3.6 jsonlite_2.0.0 dplyr_1.2.1
#> [4] compiler_4.6.0 tidyselect_1.2.1 parallel_4.6.0
#> [7] gridExtra_2.3 jquerylib_0.1.4 systemfonts_1.3.2
#> [10] scales_1.4.0 textshaping_1.0.5 yaml_2.3.12
#> [13] fastmap_1.2.0 R6_2.6.1 labeling_0.4.3
#> [16] generics_0.1.4 knitr_1.51 tibble_3.3.1
#> [19] desc_1.4.3 bslib_0.10.0 pillar_1.11.1
#> [22] RColorBrewer_1.1-3 sn_2.1.3 rlang_1.2.0
#> [25] utf8_1.2.6 cachem_1.1.0 xfun_0.57
#> [28] fs_2.1.0 sass_0.4.10 S7_0.2.2
#> [31] cli_3.6.6 pkgdown_2.2.0 withr_3.0.2
#> [34] magrittr_2.0.5 digest_0.6.39 grid_4.6.0
#> [37] lifecycle_1.0.5 vctrs_0.7.3 mnormt_2.1.2
#> [40] evaluate_1.0.5 glue_1.8.1 numDeriv_2016.8-1.1
#> [43] farver_2.1.2 ragg_1.5.2 stats4_4.6.0
#> [46] LaplacesDemon_16.1.8 rmarkdown_2.31 tools_4.6.0
#> [49] pkgconfig_2.0.3 htmltools_0.5.9