M1 · The two-stage DGP — formal specification
JoonHo Lee
2026-05-10
Source:vignettes/m1-statistical-dgp.Rmd
m1-statistical-dgp.RmdAbstract
For methodologists and reviewers who need the exact
specification behind every multisiteDGP call. This
vignette derives the joint distribution the package generates from
first principles, factors it into the four generative layers,
lists twelve testable invariants that pin the implementation to
the specification, and maps every symbol used in the package onto
the notation of the JEBS paper. You leave with a one-page
commutative diagram, a kappa-rate identity check on real simulated
output, and a citation-ready notation table.
1. What an analyst sees, what a design encodes
For a multisite trial across sites, an analyst who opens a published table sees a tuple per site:
tau_j_hat is the site’s estimated treatment effect,
se_j is its sampling standard error, n_j is
the within-site sample size, and
collects optional site-level covariates. The covariate vector is
genuinely optional: multisiteDGP’s default
formula = NULL omits it.
What the analyst does not see — but needs in order to interpret — is the latent site effect itself: the value is estimating. The sampling story says varies around with magnitude . The cross-site story says the ’s vary around a grand mean with cross-site standard deviation . The package generates both stories simultaneously, and the rest of this vignette specifies exactly how.
Concretely, a multisiteDGP simulation tibble has the
seven canonical columns shown below — every Phase 7 vignette uses the
same names.
dat_preview <- sim_multisite(preset_jebs_paper(), seed = 1L)
head(tibble::as_tibble(dat_preview)[, c(
"site_index", "z_j", "tau_j", "tau_j_hat", "se_j", "se2_j", "n_j"
)], 4)
#> # A tibble: 4 × 7
#> site_index z_j tau_j tau_j_hat se_j se2_j n_j
#> <int> <dbl> <dbl> <dbl> <dbl> <dbl> <int>
#> 1 1 -0.582 -0.116 0.652 0.426 0.182 22
#> 2 2 -0.619 -0.124 -0.315 0.577 0.333 12
#> 3 3 -1.11 -0.222 -0.633 0.256 0.0656 61
#> 4 4 1.36 0.273 0.357 0.426 0.182 22The standardized residual z_j and the latent effect
tau_j are the two latent quantities the design generates;
tau_j_hat, se_j, se2_j, and
n_j are the four observable quantities. The specification’s
job is to make every conditional and marginal in the seven-column joint
precise.
The data-generating process must therefore specify the full joint
so that a downstream user can answer at least these five questions [blueprint Ch. 4 §1]:
- What population does inform about? — needs with a defined mean and variance.
- How does precision vary across sites? — needs together with .
- What is the sampling distribution of given a site’s true effect and reported precision? — needs .
- Do covariates explain part of the latent effect? — needs .
- Is precision systematically related to the latent effect? — needs the joint structure of .
The four-layer factorization in Section 4 is one natural way to answer all five questions in a single object. Stages 1 and 2 in the next two sections are the conceptual grouping that motivates that factorization.
2. Stage 1 — the sampling distribution
Conditional on the latent site effect and on the reported sampling variance , the observed estimate is generated as a Gaussian draw:
Two points fix the convention. First, the second argument of in this package is the variance, never the standard deviation; every formula in this vignette obeys that convention. Second, the Gaussian form is justified by within-site randomization plus a central-limit argument on the difference-in-means estimator when the per-arm sample sizes are moderate and the outcome has a finite fourth moment (blueprint Ch. 4 §2.1). The package treats as known (no measurement-error layer on top of it) — this matches the standard meta-analytic convention in which reported standard errors are plugged in (Walters, 2024).
The implementation of this stage lives in gen_observations(),
which is the Layer 4 generator in the package’s four-layer
factorization. Section 4.4 below makes that connection explicit.
The unbiasedness implied by Stage 1 is the package’s testable invariant T1 (Section 7).
2.1 Stage 1 as a picture
A small simulation makes the conditional concrete. Below, 30 sites are drawn from a Gaussian-shape design with , then the observed estimates are overlaid on the latent effects on the same axis. The vertical bars give the Gaussian envelope from the Stage-1 conditional.
set.seed(1L)
design_M1 <- multisitedgp_design(
J = 30L,
sigma_tau = 0.25,
true_dist = "Gaussian",
nj_mean = 80,
cv = 0.5,
nj_min = 5,
p = 0.5,
R2 = 0,
var_outcome = 1,
dependence = "none"
)
dat_M1 <- sim_multisite(design_M1, seed = 1L)
dat_M1
#> # A multisitedgp_data: 30 sites, paradigm = "site_size"
#> # Realized vs intended:
#> # I: realized=0.519 (no target)
#> # R: realized=5.560 (no target)
#> # sigma_tau: target=0.250, realized=0.231, WARN
#> # rho_S: target=0.000, realized=0.178, PASS
#> # rho_S_marg: realized=0.178 (no target)
#> # Feasibility: WARN (n_eff=15.556)
#> # A tibble: 30 × 7
#> site_index z_j tau_j tau_j_hat se_j se2_j n_j
#> <int> <dbl> <dbl> <dbl> <dbl> <dbl> <int>
#> 1 1 -0.626 -0.157 0.680 0.348 0.121 33
#> 2 2 0.184 0.0459 0.0379 0.203 0.0412 97
#> 3 3 -0.836 -0.209 -0.0517 0.228 0.0519 77
#> 4 4 1.60 0.399 0.405 0.210 0.0440 91
#> 5 5 0.330 0.0824 -0.111 0.260 0.0678 59
#> 6 6 -0.820 -0.205 -0.168 0.199 0.0396 101
#> # ℹ 24 more rows
#> # Use summary(df) for the full diagnostic report.The print method’s diagnostic block confirms the realized between-site SD landed near the target 0.25, and that the sampling-variance product is constant across sites (this is the defining identity of the site-size-driven path — see Section 5.2).
df_stage1 <- tibble::tibble(
site = seq_len(nrow(dat_M1)),
tau_j = dat_M1$tau_j,
tau_hat_j = dat_M1$tau_j_hat,
se_j = dat_M1$se_j
)
df_stage1 <- df_stage1[order(df_stage1$tau_j), ]
df_stage1$rank <- seq_len(nrow(df_stage1))
ggplot(df_stage1, aes(y = rank)) +
geom_segment(aes(x = tau_hat_j - 1.96 * se_j,
xend = tau_hat_j + 1.96 * se_j,
yend = rank),
colour = "grey70", linewidth = 0.4) +
geom_point(aes(x = tau_j), shape = 1, size = 2, colour = "black") +
geom_point(aes(x = tau_hat_j), shape = 16, size = 1.6, colour = "#1B4965") +
geom_vline(xintercept = 0, linetype = "dashed", colour = "grey50") +
scale_y_continuous(breaks = NULL) +
labs(
x = expression(tau[j]~" / "~hat(tau)[j]),
y = "Sites (sorted by tau_j)",
title = "Stage 1 — sampling envelope around latent effect",
subtitle = "Open = tau_j (latent) · Filled = tau_j_hat (observed) · Bar = +/- 1.96 * se_j"
) +
theme_minimal(base_size = 11) +
theme(panel.grid.minor = element_blank())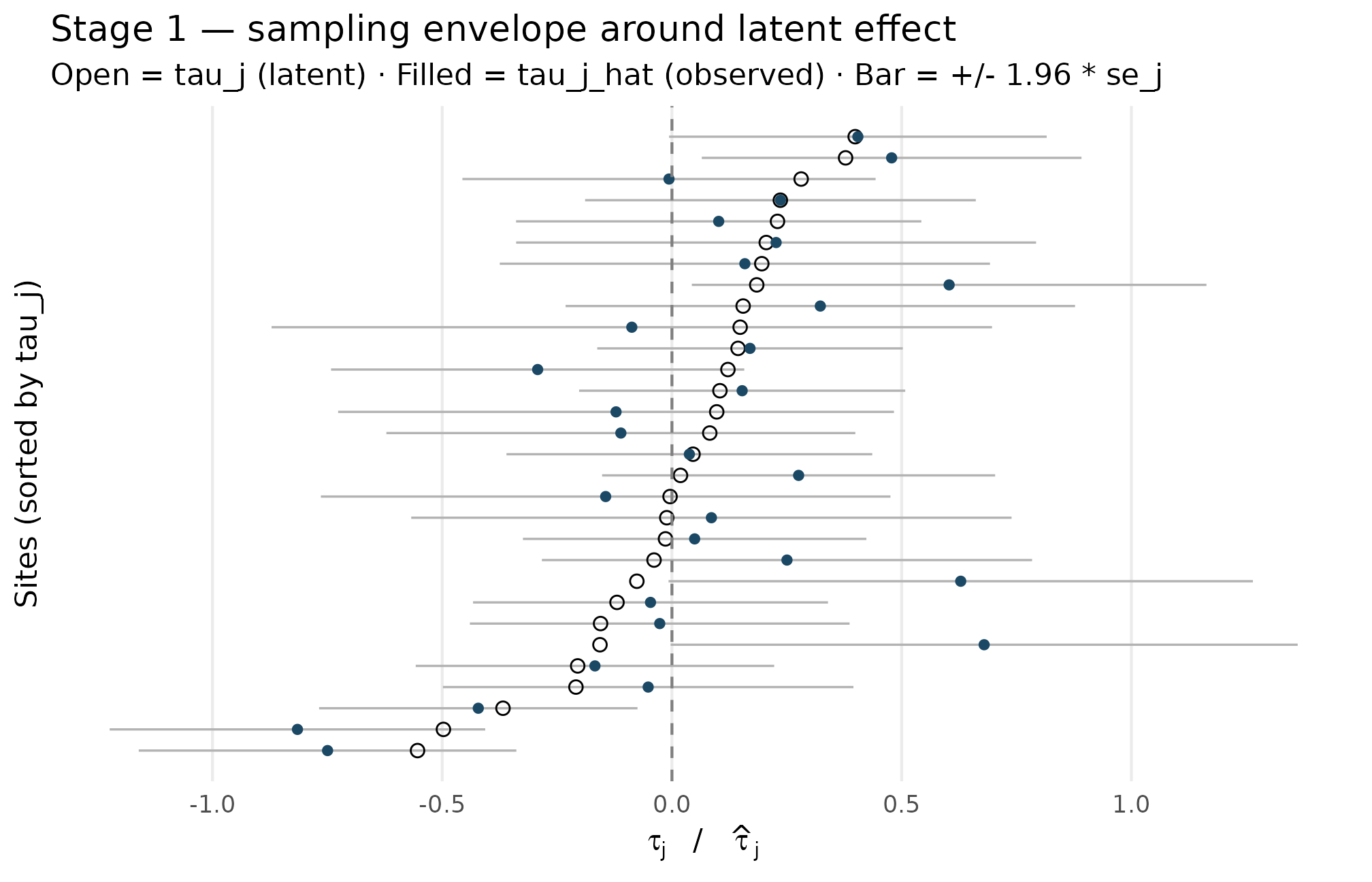
Stage 1 — observed estimates (filled dots) sit around their latent effects (open circles), with sampling envelopes scaled by site-specific se_j. Vertical spread of bar lengths reflects the precision heterogeneity from variation in n_j (range 25 to 139); the overlap between observed and latent values shrinks for the largest sites (narrowest bars). Sampling error is centered on tau_j, confirming the unbiasedness contract of T1.
Three observations to read off the figure. First, the filled dots
scatter symmetrically around the open circles — consistent with
(T1). Second, bar widths vary substantially across sites — this is
precision heterogeneity, set in Stage 2 by the site-size module (Section
4.2). Third, the bar widths are not correlated with vertical position in
the figure — consistent with the design’s
dependence = "none" choice (T7).
3. Stage 2 — the generative model on
Stage 1 takes and as inputs. Stage 2 is where they come from. The Stage-2 joint factors as a marginal on the latent effect, a precision module, and an explicit dependence coupling:
The default — and what Stage-1 unbiasedness reduces the package to in the absence of any dependence injection — is , i.e. and are independent given the design parameters [blueprint Ch. 4 §2.5]. The package allows that independence to be broken via a post-hoc rearrangement step (Section 5); this is the package’s precision dependence feature, kept optional and off by default so that the JEBS-paper specification is recovered exactly when the user does not opt in.
The next section makes the four pieces of this Stage-2 joint into four named layers and gives each one its conditional distribution.
4. The four generative layers
The full joint, written across all of Stages 1 and 2, factors into four layers in a fixed order: latent effects → site-size margins → precision dependence → observation draws. Each layer corresponds to one named generator family in the package, and one section of this vignette below.
The order is operational: Layer 1 fills z_j and
tau_j, Layer 2 fills n_j and
se2_j (and therefore se_j), Layer 3 (when
active) permutes the precisions to induce a target rank correlation with
the effects, and Layer 4 fills tau_j_hat. The four layers
are also the four S3 generators gen_effects(), gen_site_sizes(),
align_rank_corr()
(and its copula and hybrid siblings), and gen_observations().
4.1 Layer 1 — latent effects
Site labels carry no information by design — they are interchangeable. de Finetti’s exchangeability theorem then implies that the latent effects are conditionally i.i.d. from some mixing distribution :
This corresponds to JEBS Equation (2) when is taken to be Gaussian (Lee et al., 2025). The package implements eight choices for — Gaussian, two-component mixture, asymmetric Laplace (Yu & Zhang, 2005), Student-t, skew-normal (Azzalini, 1985), point-mass-plus-slab, a user-supplied callback, and a Dirichlet process mixture stub. The catalog and standardization details live in M2: G-distribution catalog; for the purposes of M1, the only contract that matters is that every member of the catalog has mean and variance on the output scale.
The package factors as a location plus a scaled standardized residual:
where
is the standardized version of the chosen
shape. The column z_j in the simulation tibble is exactly
this standardized residual. Var
is the package’s testable invariant T2 (Section 7), and writing the
latent effect this way is what makes the covariate-augmented form
a transparent extension rather than a reparameterization. Under
covariates,
is the residual between-site SD — the marginal SD also picks up
.
This is why print.multisitedgp_data()
shows two
values when covariates are active. The covariate-augmented case is the
subject of A4:
Covariates and precision dependence; the covariate-off identity
is the package’s testable invariant T6.
4.2 Layer 2 — site-size margins
Two pieces. First, site sample sizes are drawn from a truncated Gamma distribution moment-matched to the user-supplied mean and coefficient of variation , with a hard lower bound at [blueprint Ch. 4 §2.4]:
The default is enforced as the package’s testable invariant T10 (Section 7); the truncation is the reason no multisite simulation ever produces an smaller than 5 by default. The truncated-Gamma form is the modern Engine A2; the legacy JEBS-appendix Engine A1 uses a censor-and-round form instead, and the two engines exist precisely so that the JEBS appendix is reproducible bit-for-bit (T1a) without giving up modern moment-matching (T1b). See M3: Margin and SE models for the engine contracts.
Second, conditional on , the per-site sampling variance is set deterministically:
with the proportionality constant derived from a balanced two-arm randomization with allocation share , residual outcome variance , and covariate-explained share :
At the package defaults , , , this collapses to . The deterministic form is what makes the identity hold for every site in every simulation under the site-size-driven path — this is the package’s testable invariant T3 (Section 7) and the defining contract of the path itself. A minute of code makes it visible:
range(dat_M1$se2_j * dat_M1$n_j)
#> [1] 4 4
unique(round(dat_M1$se2_j * dat_M1$n_j, 8))
#> [1] 4
compute_kappa(p = 0.5, R2 = 0, var_outcome = 1)
#> [1] 4The first line shows the kappa-rate product is constant. The third line confirms the closed form returns 4.
The alternative direct-precision path
(gen_se_direct()) discards the deterministic kappa identity
in exchange for direct control of the informativeness scalar
and the precision heterogeneity ratio
;
that path is the subject of M3 §3 and is outside the scope of this
vignette except as part of the JEBS notation table in Section 6.
4.3 Layer 3 — precision dependence
By default, and are independent because they are generated from independent random draws. When a user asks for a non-zero target Spearman or Pearson correlation, the package performs a post-hoc rearrangement of the precisions so that their multiset is preserved exactly while their pairing with effects matches the target.
Three coupling functions are exposed: align_rank_corr()
(rank permutation), align_copula_corr()
(Gaussian copula re-pairing), and align_hybrid_corr()
(copula initialization plus rank refinement). The methodological
contrast across the three lives in M4: Precision dependence
theory.
The contracts that matter for M1 are:
- When
dependence = "none", the coupling reduces to the identity permutation (T7). No rearrangement, no change to either marginal. - The marginal of is preserved exactly (its mean, variance, and shape are not touched).
- The multiset of is preserved exactly (its realized values are not touched, only their pairing with sites is).
- The realized informativeness is preserved exactly across any active rearrangement.
The reason the package generates first and rearranges second — rather than building dependence into a joint distribution upfront — is to keep the user’s specification of the marginals from drifting. T7 is the testable contract that the rearrangement is genuinely identity when the user opts out.
4.4 Layer 4 — observation draws
Layer 4 is the implementation of Stage 1 from Section 2: given the latent effects produced by Layer 1 and the sampling variances produced by Layer 2 (possibly rearranged in Layer 3), draw
independently across
.
The implementation is gen_observations();
the column tau_j_hat in the simulation tibble is its
output.
Independence of the per-site draws (conditional on Layer 1–3 outputs) is part of the same blueprint Ch. 4 §2.5 default. It is not a separate testable invariant — it is the defining form of the observation generator. T1 in Section 7 covers the unbiasedness contract; the conditional independence of the draws is structural.
4.5 The Stage-2 joint as a picture
The default joint — independent marginals, no dependence coupling — looks like this. The two density panels show the marginal of each axis, and the central scatter shows the realized pairs.
set.seed(2L)
design_marg <- multisitedgp_design(
J = 200L,
sigma_tau = 0.25,
true_dist = "Gaussian",
nj_mean = 60,
cv = 0.5,
nj_min = 5,
p = 0.5,
R2 = 0,
var_outcome = 1,
dependence = "none"
)
dat_marg <- sim_multisite(design_marg, seed = 2L)
sd_tau <- round(sd(dat_marg$tau_j), 4)
gm_se2 <- round(exp(mean(log(dat_marg$se2_j))), 4)
range_se2 <- range(dat_marg$se2_j)
sd_tau; gm_se2; range_se2
#> [1] 0.2685
#> [1] 0.0791
#> [1] 0.01960784 0.50000000
df_marg <- tibble::tibble(
tau_j = dat_marg$tau_j,
se2_j = dat_marg$se2_j
)
p_center <- ggplot(df_marg, aes(x = tau_j, y = se2_j)) +
geom_point(alpha = 0.45, size = 1.6, colour = "#1B4965") +
geom_vline(xintercept = 0, linetype = "dashed", colour = "grey50") +
labs(
x = expression(tau[j]~" (latent effect)"),
y = expression(widehat(se)[j]^{2}~" (sampling variance)")
) +
theme_minimal(base_size = 11) +
theme(panel.grid.minor = element_blank())
p_top <- ggplot(df_marg, aes(x = tau_j)) +
geom_density(fill = "#1B4965", alpha = 0.35, colour = "#1B4965") +
labs(x = NULL, y = "density") +
theme_minimal(base_size = 9) +
theme(panel.grid.minor = element_blank(),
axis.text.x = element_blank(),
axis.ticks.x = element_blank())
p_right <- ggplot(df_marg, aes(x = se2_j)) +
geom_density(fill = "#62929E", alpha = 0.45, colour = "#62929E") +
coord_flip() +
labs(x = NULL, y = "density") +
theme_minimal(base_size = 9) +
theme(panel.grid.minor = element_blank(),
axis.text.y = element_blank(),
axis.ticks.y = element_blank())
# Compose without depending on patchwork (soft dep).
gridExtra::grid.arrange(
p_top, grid::nullGrob(),
p_center, p_right,
ncol = 2, widths = c(4, 1), heights = c(1, 4)
)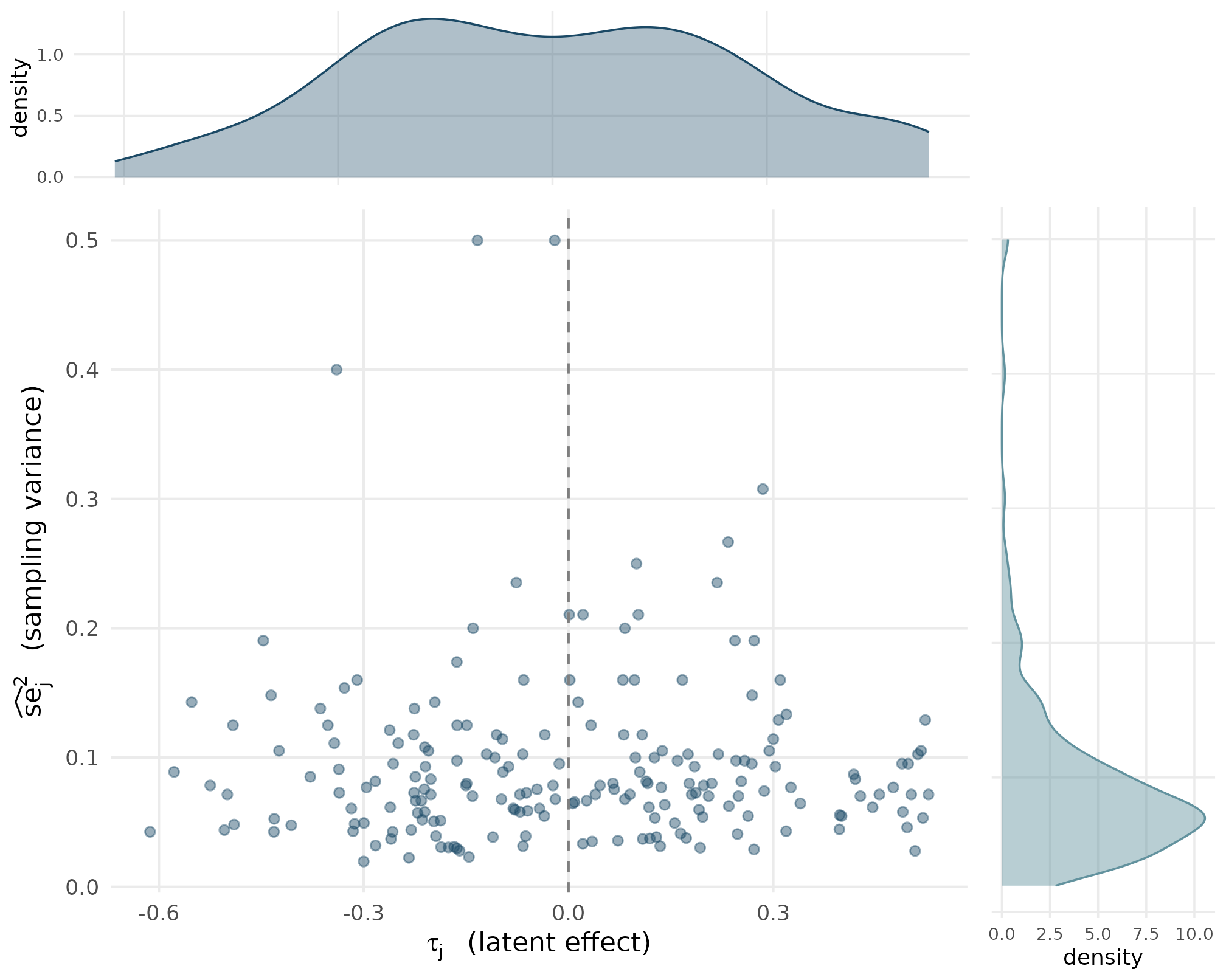
Stage 2 default joint p(tau_j, se_j squared) at J = 200. Top: marginal density of latent effects, centered near 0 with SD near sigma_tau = 0.25 (T2). Right: marginal density of sampling variances, right-skewed because n_j is right-skewed and se squared = kappa over n_j (T3). Center: scatter of realized (tau_j, se squared) pairs is roughly cloud-shaped, with no visible tilt — independence at the sample level, consistent with T7.
Three things to read off the joint:
- The top panel’s density is centered near zero and has spread close to . This is T2 in action: the standardized residual has unit variance, so ’s marginal SD lands at .
- The right panel’s density is right-skewed. That is the deterministic consequence of (T3) plus the truncated-Gamma marginal on : large pile up near the mean, but small pull the SE-squared distribution out to the right.
- The central scatter is roughly cloud-shaped. There is no systematic tilt — the realized rank correlation between and is statistically indistinguishable from zero at . This is T7 in action.
5. Default-mode reduction
When every option is at its default value, the package reduces to a
short, vanilla two-stage random-effects model. Defaults are
true_dist = "Gaussian", formula = NULL,
dependence = "none", the site-size-driven path
(Paradigm A in the blueprint), and
,
,
giving
:
This is the site-size-driven path’s default form; the JEBS-paper specification is contained inside it as a special case (Lee et al., 2025). Section 6 maps every symbol to JEBS notation, and Section 7 lists the twelve invariants the package carries.
5.1 Two paths through Stage 2
The blueprint distinguishes two ways to set the precision marginal. The site-size-driven path (Paradigm A in the blueprint) is what Section 4.2 derived: a user supplies , , and ; sites sizes are drawn; precisions follow as . The direct-precision path (Paradigm B in the blueprint) inverts: a user supplies the informativeness scalar and the precision heterogeneity ratio , and the precision marginal is reconstructed (log-evenly spaced) to match. The default is the site-size-driven path; the direct-precision path is the subject of M3: Margin and SE models. The notation table in Section 6 covers symbols from both paths.
5.2 Three deviations from the default
Three knobs, each with a default that recovers the JEBS-paper specification, expand the package beyond the JEBS scope:
-
Covariate-augmented Stage 2 — passing a
formulawithdataandbetaintroduces the linear term (Section 4.1). Withformula = NULL, the unconditional and covariate-augmented code paths are identical to floating-point precision (T5). -
Precision dependence — setting
dependencetorank,copula, orhybridactivates Layer 3’s rearrangement (Section 4.3). Withdependence = "none", the rearrangement reduces to identity (T7). -
G-shape choice — setting
true_distto one of the seven non-Gaussian shapes (Section 4.1) selects from M2’s catalog. Withtrue_dist = "Gaussian", the latent effects follow JEBS Equation- exactly.
6. JEBS notation map
This section maps every symbol the package uses onto the notation of the JEBS paper (Lee et al., 2025). The table is split into three parts: equations from the body of the JEBS paper, simulation-design formulas from the JEBS appendix code, and package-only extensions that have no JEBS counterpart.
6.1 JEBS paper body equations
| JEBS reference | Symbol / equation | Package mapping | Notes |
|---|---|---|---|
| Stage-1 setup (unnumbered) | gen_observations() |
Justified by within-site randomization + CLT | |
| Eq. (1) |
compute_kappa() (closed form) |
Per-site SE under balanced randomization | |
| Eq. (2) |
gen_effects() with
true_dist = "Gaussian"
|
Default G shape; mean , variance | |
| Eq. (3) | Posterior shrinkage estimator | (out of DGP scope) | Estimator side; covered in siteBayes2, not
multisiteDGP
|
| Eq. (4) | informativeness() |
Generative-side diagnostic |
6.2 JEBS appendix simulation formulas
| Appendix reference | Symbol / construct | Package mapping | Notes |
|---|---|---|---|
| Appendix mixture form | Two-component mixture with parameters |
gen_effects() with
true_dist = "Mixture"
|
Bit-identical with appendix code under Engine A1 (T1a) |
| Appendix site-size + SE form | Gamma + censor + round; |
gen_site_sizes()
Engine A1 |
T1a fixture |
| Appendix observation form | + row shuffle |
gen_observations()
Engine A1 path |
Order of RNG calls is itself an invariant (T12) |
| Appendix default | , , → |
multisitedgp_design()
defaults |
T2/T3 fixture |
| Appendix G shapes | Gaussian, two-component mixture, asymmetric Laplace |
gen_effects()
true_dist ∈ {Gaussian, Mixture, ALD} |
Three of the package’s eight shapes |
| Appendix site-size marginal |
gen_site_sizes()
Engine A2 |
T10 fixture |
6.3 Package-only extensions
| Feature | Symbol / construct | Package mapping | Notes |
|---|---|---|---|
| Site-level covariates |
gen_effects() with
formula, beta, data
|
Reduces to JEBS Eq. (2) when formula = NULL (T5) |
|
| Precision dependence (rank) | Target Spearman between (or ) and | align_rank_corr() |
Identity permutation when dependence = "none" (T7) |
| Precision dependence (copula) | Target Pearson via Gaussian copula | align_copula_corr() |
M4 vignette covers selection criteria |
| Precision dependence (hybrid) | Copula initialization + rank refinement | align_hybrid_corr() |
M4 vignette covers when each path is appropriate |
| Direct-precision path | Inverse design from to marginal | gen_se_direct() |
M3 vignette covers feasibility region |
| Five new G shapes | Student-t, skew-normal (Azzalini, 1985), point-mass-plus-slab, user, DPM stub |
gen_effects()
true_dist ∈ {StudentT, SkewN, PointMassSlab, User,
DPM} |
Five of the package’s eight shapes; Gaussian default
preserves JEBS Eq. (2) |
7. Twelve testable invariants
The specification in Sections 1–4 is only as precise as the
implementation it pins down. Twelve invariants, each a one-line contract
on observable output, form the testable backbone of this correspondence
— every one of them lives in
tests/testthat/test-jebs-parity.R and surrounding test
files in the package source. The numbering follows the blueprint Ch. 4
§7 listing, with T1 split into T1a (Engine A1, bit-identical with the
JEBS appendix) and T1b (Engine A2, distributional parity) per the
blueprint’s R1.4 fix.
T1a — Engine A1 bit-identical parity (mixture)
Engine A1 (legacy, legacy_jebs_censor_round) under the
JEBS appendix’s exact configuration — true_dist = "Mixture"
with
,
,
,
,
,
formula = NULL, dependence = "none" — produces
a simulation tibble whose xxhash64 digest matches the digest of the JEBS
appendix pipeline output across the seed set
.
Bit-identical means floating-point byte equality, not
distributional similarity.
T1b — Engine A2 distributional parity (Gaussian)
Engine A2 (modern, trunc_gamma_moment) with
true_dist = "Gaussian" and the package defaults
(,
,
,
,
,
,
,
)
produces
samples whose empirical mean is within
of
,
whose empirical variance is within
of
,
and whose KS test against
passes at the 0.05 level in at least 95 percent of
Monte Carlo replications. Distributional parity, not
bit-identical.
T2 — kappa is exactly 4 at defaults
At , , , the realized equals 4 exactly (no rounding). This is the bridge from the closed form to the package output the user sees. Verified directly:
compute_kappa(p = 0.5, R2 = 0, var_outcome = 1)
#> [1] 4
identical(compute_kappa(p = 0.5, R2 = 0, var_outcome = 1), 4)
#> [1] TRUET3 — kappa-rate identity holds for every site
Under the site-size-driven path,
for every
to floating-point precision. Equivalently, the within-design variance of
across
is zero. The check on dat_M1 from Section 4.2 is the
operational form of this contract.
T4 — Stage-2 independence under
dependence = "none"
When dependence = "none", the realized Spearman and
Pearson correlations between
and
are statistically indistinguishable from zero. The acceptance band is
the empirical 95th percentile of
from 1000 random independent permutations (a permutation
envelope) — not the Fisher-z asymptotic critical value, which is a
Pearson approximation that is tight but inexact for Spearman. The
blueprint corrected an earlier draft on this point; the package follows
the permutation form.
T5 — covariate-off equivalence
When formula = NULL (or when the user passes
beta = 0), the covariate-augmented code path produces
output identical to the unconditional path to floating-point precision
().
This is what makes the package’s covariate handling a strict extension
rather than a reparameterization.
T6 — residual equals marginal SD when no covariates
When formula = NULL, the
print.multisitedgp_data() method reports
sigma_tau_resid and sigma_tau_marg as
numerically identical, and both equal the input sigma_tau
to floating-point precision. With covariates, the marginal SD picks up
and the two quantities diverge — see A4.
T7 — precision dependence reduces to identity
When dependence = "none", the alignment functions align_rank_corr(),
align_copula_corr(),
and align_hybrid_corr()
all reduce to the identity permutation: input vector equals output
vector, exactly. This is the operational form of the “off by default”
contract on Layer 3.
T8 — informativeness formula
informativeness()
implements JEBS Equation (4) verbatim:
where the bar denotes the arithmetic mean over . Section 6.1 entry for Eq. (4) is the same statement.
T9 — default G shape is Gaussian
When true_dist is omitted, the package defaults to
"Gaussian". At
across
Monte Carlo replications, the Shapiro-Wilk test against
rejects at the 0.01 level a count consistent with the binomial
acceptance interval — concretely, the count of
events falls in
for
.
The acceptance band acknowledges that under a true normal, the
Shapiro-Wilk
-value
is uniform, so the count of small
-values
is itself a binomial random variable.
T10 — site-size truncation enforces
At , no ever appears in the simulation output, and the realized mean is within percent of the requested mean. The truncation is hard, not soft, in both engines.
T11 — informativeness range matches JEBS over the design grid
Across the JEBS simulation grid — , , , giving design cells — the realized informativeness values cover the JEBS-reported range (Lee et al. 2025, p. 13). The full grid is the contract; a default-CV slice is not sufficient.
T12 — Engine A1 RNG sequence is locked
Under Engine A1, the random-number-generator call sequence is
exactly: rgamma(J) for site sizes, rnorm(J)
for the first noise draw, then the row-shuffle internal
sample() call from
dplyr::slice_sample(prop = 1), then rnorm(J)
for any subsequent noise. No additional RNG calls. Reordering, omitting
the slice_sample shuffle, or re-seeding mid-pipeline all
violate T12 and break T1a parity simultaneously. Engine A2’s RNG
sequence is separate and is the subject of T20 in the blueprint (out of
M1 scope).
How the invariants relate to the four layers
A short cross-reference, for orientation:
| Invariant | Layer | What it pins |
|---|---|---|
| T1, T1a, T1b | Layer 4 (and Engine A1/A2 pipelines end-to-end) | Stage 1 unbiasedness; appendix bit-equivalence; modern distributional parity |
| T2, T3 | Layer 2 | at defaults; deterministic kappa-rate identity |
| T4, T7 | Layer 3 | Independence under dependence = "none"; identity
permutation contract |
| T5, T6 | Layer 1 | Covariate-off equivalence; residual-equals-marginal SD |
| T8, T11 | Cross-layer (diagnostics) | Informativeness formula; informativeness range |
| T9 | Layer 1 | Default G shape is Gaussian |
| T10 | Layer 2 | Site-size truncation |
| T12 | Engine A1 | RNG call order locked to JEBS appendix |
Forward pointers — T13–T20 in companion vignettes
T1–T12 above are the invariants that fit naturally inside the two-stage DGP specification this vignette covers. The blueprint’s full T-numbered list extends further; the remaining invariants live with the layer they constrain:
- T13–T17 — per-shape unit-variance standardization — checks that ALD, Mixture, Student-, skew-normal, and point-mass-slab generators all return at the package’s closed-form rescalers. Operationalized in M2: G-distribution catalog and standardization §4 (sanity checks) and the package’s test suite.
- T18–T19 — covariate decomposition + Spearman tie handling — identifiability of the residual / marginal split when covariates enter Layer 1, and the rank-correlation behavior at tied precision values. Operationalized in A4: Covariates and precision dependence for the residual-vs-marginal demonstration and in M4: Precision dependence theory §3.2 for the rank ties.
- T20 — Engine A2 reproducibility — the modern-engine RNG-call sequence (the analogue of T12 for Engine A2) and its hash determinism across reruns. Operationalized in M7: Reproducibility and provenance §3 (canonical-hash determinism) and §5 (manifest pattern).
8. Limitations and assumptions
The two-stage specification carries seven assumptions that are explicit in blueprint Ch. 4 §8 and worth surfacing here:
-
Independence baseline. By default,
.
Activating
dependencebreaks this on purpose (Section 4.3, M4). - Linear covariate effect. The covariate term enters as . Nonlinear relationships are reachable only via a user-supplied callback (M5).
- Finite second moments of . is required. Heavy-tailed Student-t with is rejected at construction.
- Within-site Normality of the per-site estimate. CLT-justified for moderately large; for very small sites () the approximation is tighter but degrades.
- Effect-size scale. is the default; changing it rescales multiplicatively.
- Single treatment arm with complete randomization within site. Cluster-randomized, blocked, and factorial designs are outside the current package surface.
- No measurement error in . The package treats reported standard errors as exact, matching the standard meta-analytic plug-in convention (Walters, 2024).
9. Where to next
- M2: G-distribution catalog and standardization — the eight choices for , their parameterizations, and how standardization is enforced (T2 is operationalized there).
- M3: Margin and SE models — Engine A1 vs. Engine A2, and the direct-precision path.
- M4: Precision dependence theory — the three injection methods and when each is appropriate.
- M7: Reproducibility and provenance — canonical hashes, RNG-state capture, and how to write a defensible methods appendix.
- For a worked applied example that touches every layer, A1: Getting started; for the diagnostic rubric that turns the invariants in Section 7 into PASS / WARN / FAIL flags on a single simulation, A3: Diagnostics in practice.
References
The foundational data-generating process and the JEBS preset implement the design in Lee et al. (2025). Empirical-Bayes context for site-effect estimation, including the assumption that reported sampling standard errors are taken as known, is collected in Walters (2024). The asymmetric Laplace shape used in the catalog is from Yu & Zhang (2005), and the skew-normal shape is from Azzalini (1985). The applied calibration benchmarks underlying the education presets are from Weiss et al. (2017).
The Stage-1 Gaussian-sampling justification (CLT-based on within-site randomization with finite fourth moments) is the standard meta-analytic baseline; the package’s blueprint Ch. 4 §2.1 gives the formal motivation. Specific canonical references in the meta-analytic methodology literature are left to the analyst’s choice rather than prescribed here.
Acknowledgments
This research was supported by the Institute of Education Sciences, U.S. Department of Education, through Grant R305D240078 to the University of Alabama. The opinions expressed are those of the authors and do not represent views of the Institute or the U.S. Department of Education.
Session info
#> R version 4.6.0 (2026-04-24)
#> Platform: x86_64-pc-linux-gnu
#> Running under: Ubuntu 24.04.4 LTS
#>
#> Matrix products: default
#> BLAS: /usr/lib/x86_64-linux-gnu/openblas-pthread/libblas.so.3
#> LAPACK: /usr/lib/x86_64-linux-gnu/openblas-pthread/libopenblasp-r0.3.26.so; LAPACK version 3.12.0
#>
#> locale:
#> [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8
#> [4] LC_COLLATE=C.UTF-8 LC_MONETARY=C.UTF-8 LC_MESSAGES=C.UTF-8
#> [7] LC_PAPER=C.UTF-8 LC_NAME=C LC_ADDRESS=C
#> [10] LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
#>
#> time zone: UTC
#> tzcode source: system (glibc)
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ggplot2_4.0.3 multisiteDGP_0.1.1
#>
#> loaded via a namespace (and not attached):
#> [1] gtable_0.3.6 jsonlite_2.0.0 dplyr_1.2.1 compiler_4.6.0
#> [5] tidyselect_1.2.1 nleqslv_3.3.7 gridExtra_2.3 jquerylib_0.1.4
#> [9] systemfonts_1.3.2 scales_1.4.0 textshaping_1.0.5 yaml_2.3.12
#> [13] fastmap_1.2.0 R6_2.6.1 labeling_0.4.3 generics_0.1.4
#> [17] knitr_1.51 tibble_3.3.1 desc_1.4.3 bslib_0.10.0
#> [21] pillar_1.11.1 RColorBrewer_1.1-3 rlang_1.2.0 cachem_1.1.0
#> [25] xfun_0.57 fs_2.1.0 sass_0.4.10 S7_0.2.2
#> [29] cli_3.6.6 pkgdown_2.2.0 withr_3.0.2 magrittr_2.0.5
#> [33] digest_0.6.39 grid_4.6.0 lifecycle_1.0.5 vctrs_0.7.3
#> [37] evaluate_1.0.5 glue_1.8.1 farver_2.1.2 ragg_1.5.2
#> [41] rmarkdown_2.31 tools_4.6.0 pkgconfig_2.0.3 htmltools_0.5.9